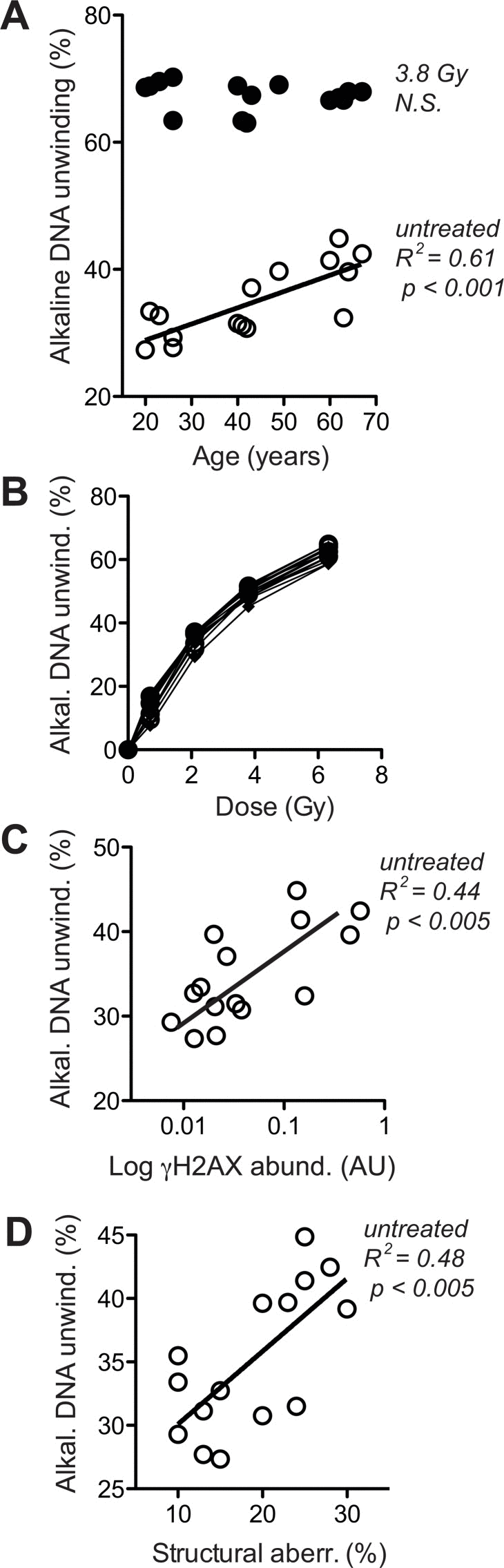
Figure 5.Levels of DNA strand breaks at base line and following IR exposure(A) Fluorimetric quantitation of alkaline DNA unwinding carried out with lysates of untreated cells (open circles) and cells exposed to 3.8 Gy (closed circles). Values are normalised to total signal intensity corresponding to non-unwound DNA. (B) IR-dose-response curves determined as in (A). (C) Cross-comparison of base line H2AX phosphorylation (in the absence of VP16, same data as represented by open symbols in Fig. 4A) and base line alkaline DNA unwinding in the absence of IR (same data as open circles in section A of this figure). (D) Cross-comparison of structural chromosome aberrations (same data as in Fig. 3D) and base line alkaline DNA unwinding in the absence of IR (same data as open circles in section A of this figure). (A, C, D) Results of linear regression are stated as R2: Pearson's coefficient for goodness of fit, p: probability for slope = 0, and N.S.: no significant correlation. Data represent mean values of five independent cell cultures per donor, each analysed in four technical replicates. SEM was in all cases less than 20% of the values and is omitted for the sake of clarity.