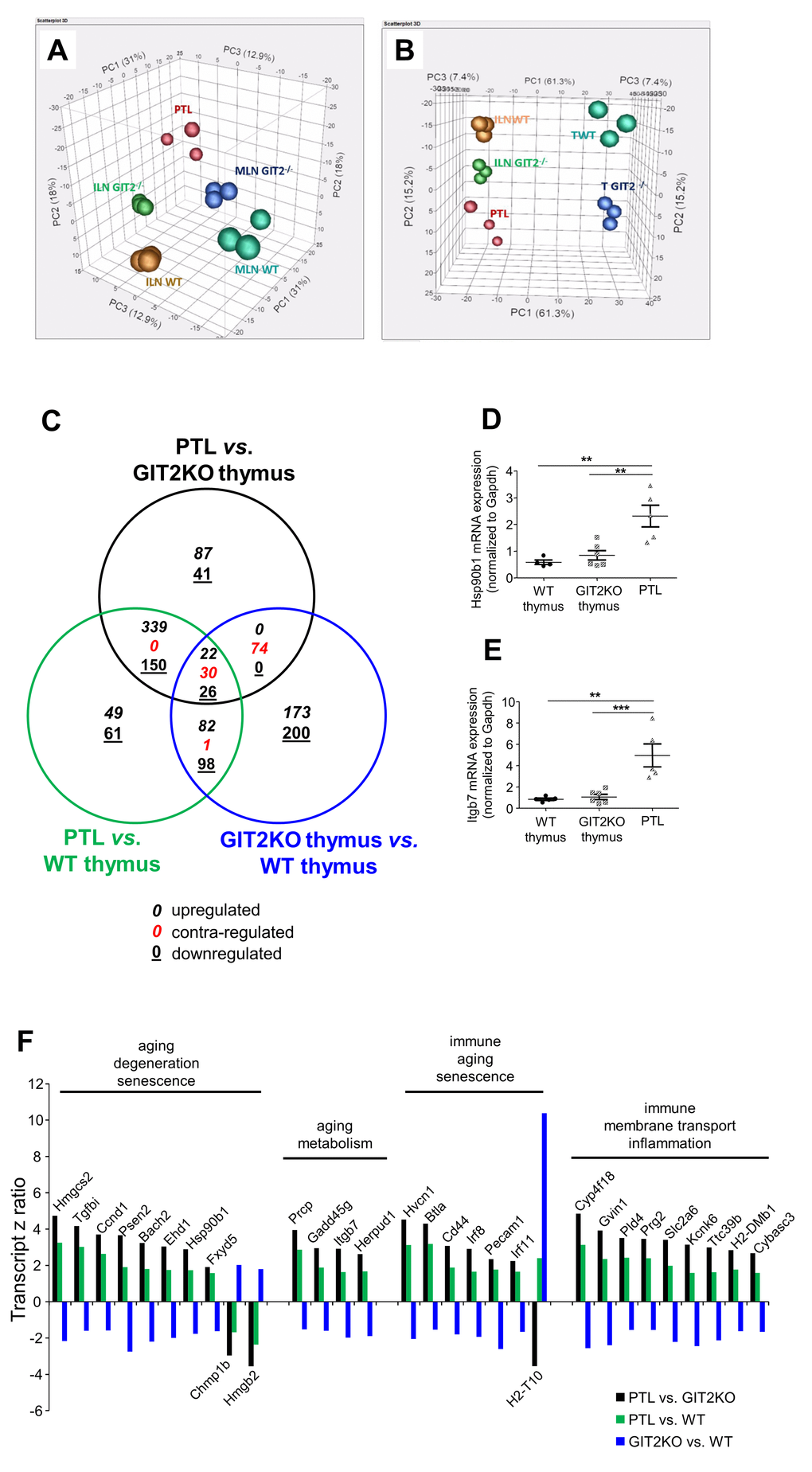
Figure 4.Transcriptomic analysis of GIT2KO parathymic lobes. Principal component analyses were performed upon transcriptomic data from thymus (WT thymus = TWT; GIT2KO thymus = T GIT2-/-), WT/GIT2KO ILN and MLN, as well as PTLs from GIT2KO mice. PTL transcriptome data separates with PCA from ILN and MLN from both WT and GIT2KO mice (A). PTLs share component 2 with GIT2KO thymi and component 1 with WT and GIT2KO ILN (B). Venn diagram analysis of significantly-regulated transcripts generated by the following tissue transcriptome comparisons: GIT2KO PTLs vs. GIT2KO thymus (black circle); GIT2KO PTL vs. WT thymus (green circle); GIT2KO thymus vs. WT thymus (blue circle). For the Venn diagram numbers in italics represent upregulated transcripts, underlined numbers represent downregulated transcripts, red numbers represent transcripts possessing diverse expression polarities (C). RT-PCR validations of selected PTL transcripts Hsp90β1 (D) and Itgb7 (E: **p<0.01, ***p<0.001). Transcriptomic z ratios of 30 common and coherently regulated (same expression polarity across three Venn sectors) transcripts generated from Venn separation of PTL-associated transcripts (F).