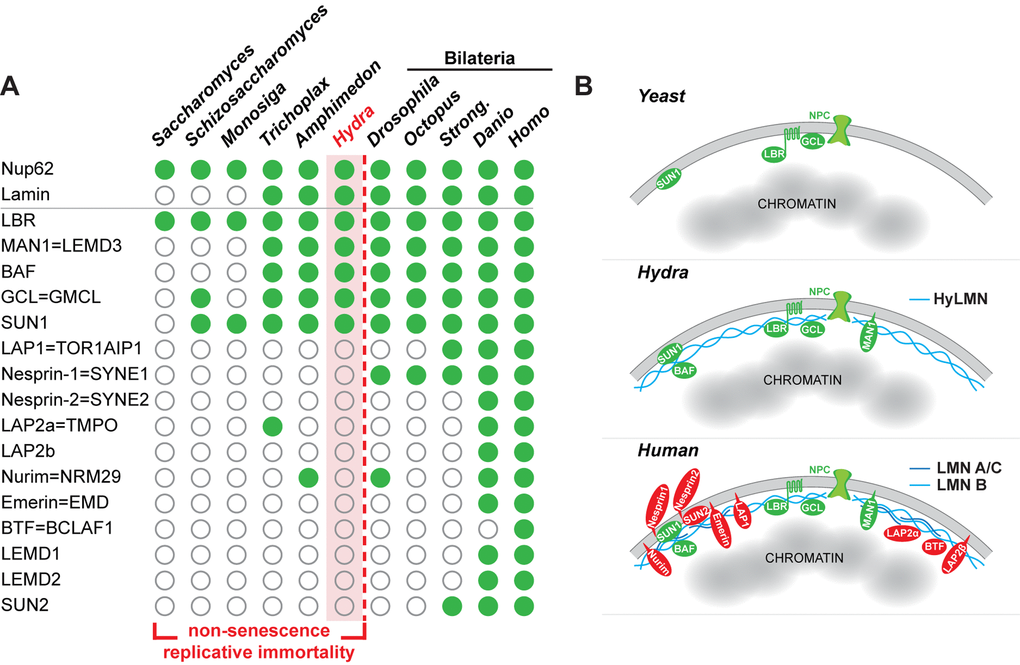
Figure 11.The complexity of the nuclear lamina architecture in Hydra and other pre-bilaterian animals resembles that of the unicellular eukaryotes. (A) Phylogenetic distribution of genes coding for Lamin and lamin-binding proteins (rows) as revealed by a BLAST-screening of genomes (columns) from unicellular and multicellular eukaryotes. Filled green circles indicate genes identified with high confidence, open circles indicate genes not found or identified with low confidence. Raw data including e-values and sequence accession numbers are presented in the Supplementary Dataset 1. Nuclear pore protein Nup62 used as a control demonstrates ubiquitous presence in eukaryotic organisms. The repertoire of genes coding for lamin-binding proteins in Hydra is essentially similar to that of a placozoan Trichoplax, sponge Amphimedon, and the unicellular organisms - Sacchoromyces, Schizosaccharomyces and Monosiga. We suggest that low complexity of the nuclear envelope in these animals enables for their non-senescence and replicative immortality. Members of the Bilateria are characterized by a rich repertoire of lamin-binding proteins and higher complexity of the nuclear envelope. Somatic cells of these animals lose replicative immortality, ultimately leading to a restricted lifespan. (B) A model representing increasing complexity of the nuclear envelope architecture in eukaryotes - from yeast and pre-bilaterian Metazoa, as Hydra, to human. Conserved proteins are in green, lamin-binding proteins that are present only in higher bilaterian animals are highlighted in red.