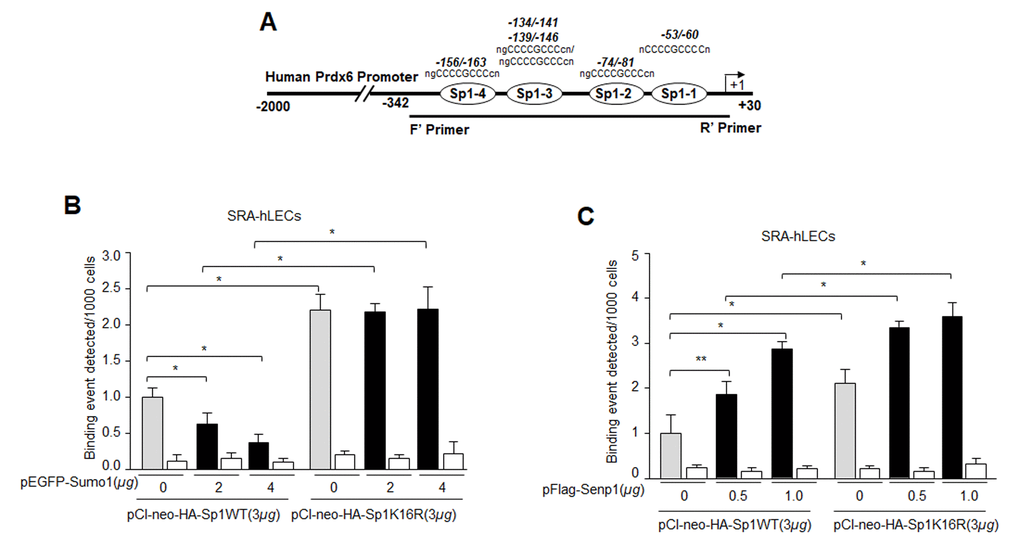
Figure 8.Mutagenesis and in vivo DNA binding assay revealed increased binding of Sumoylation-deficient Sp1K16R to Sp1 site in Prdx6 promoter by skipping aberrant Sumoylation effect. (A) Schematic representation of the regulatory region of proximal promoter of Prdx6 gene containing GC-box (Sp1 binding sites) showing primer location used in ChIP assay. (B) Sumo1 failed to affect Sp1-DNA binding activity in Sumoylation-deficient Sp1K16R transfected LECs. SRA-hLECs were transfected with either pCl-neo-HA-Sp1 or its mutant pCl-neo-HA-Sp1K16R alone or cotransfected with different concentrations of pEGFP-Sumo1. ChIP experiment was carried out as described in Materials and Methods. Chromatin samples prepared from LECs cotransfected with pEGFP-Sumo1 with either pCl-neo-HA-Sp1 or its mutant pCl-neo-HA-Sp1K16R were subjected to ChIP assay with a ChIP grade antibody, anti-HA (gray and black bars) and control IgG (open bars). The DNA fragments were used as templates for RT-qPCR by using primers designed to amplify −342 to +30 region of the Prdx6 gene promoter bearing Sp1 binding sites as shown. Histogram shows the amplified DNA through real-time qPCR analysis. 0 µg vs 2 µg and 4µg pEGFP-Sumo1, pCl-neo-HA-Sp1 WT vs pCl-neo-HA-Sp1K16R (*p < 0.001). (C) Senp1 overexpression showed increased Sp1 DNA binding of Sp1WT and comparable to Sumoylation deficient Sp1K16R. SRA-hLECs were cotransfected with pFlag-Senp1 with either pCl-neo-HA-Sp1 or mutant pCl-neo-HA-Sp1K16R as indicated. ChIP assay was conducted as described above using anti-HA antibody. Histograms represent the concentration dependence of Senp1-induced enrichment of Sp1 at its binding sites in Prdx6 gene promoter. 0 µg vs 0.5 µg and 1µg pFlag-Senp1, pCl-neo-HA-Sp1 WT vs pCl-neo-HA-Sp1K16R (**p<0.05; *p < 0.001).