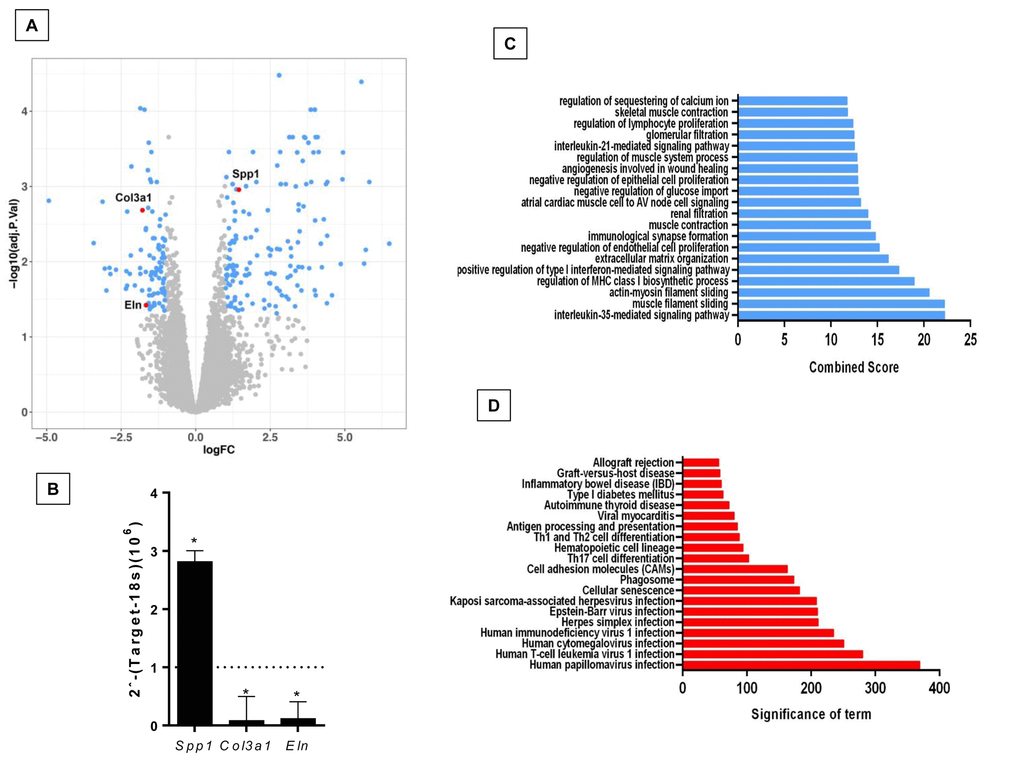
Figure 2.Differentially expressed genes and pathways in lungs from old compared to young Zmpste24 deficient mice. (A) Volcano plot of the global gene expression profiling in lungs from old Zmpste24 deficient mice vs young littermates. Each point represents the difference in expression (log fold-change) between the two groups of mice plotted against the level of statistical significance. Right blue dots represent overexpressed genes; left blue dots represent downregulated genes at a significant level of p< 0.05. Some Core Matrisome genes are plotted in red. (B) The expression of these genes was evaluated by quantitative RT-PCR. Data are shown as fold-change values as compared with young -/- (dotted lines). (C and D) Gene ontology (C) and KEGG (D) functional analysis of dysregulated genes in old Zmpste24 deficient mice compared to young littermates. The most significant 20 terms are shown in this figure. Threshold criteria considered for the analysis are log Fold change > 1 or <-1 and p-value < 0.05.