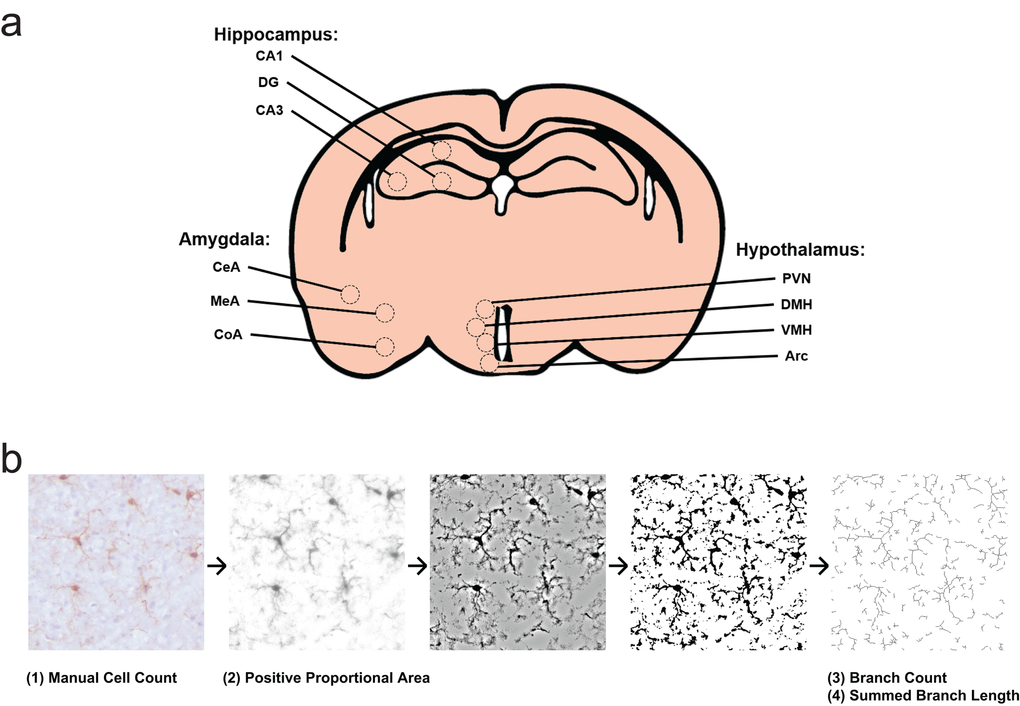
Figure 4.Experimental design for morphology survey. (a) Select subfields and nuclei analyzed within the hippocampus, hypothalamus, and amygdala. CA1, cornu ammonis 1; CA3, cornu ammonis 3; DG, dentate gyrus; Arc, arcuate nucleus; DMH, dorsomedial hypothalamic nucleus; PVN, paraventricular nucleus; VMH, ventromedial hypothalamic nucleus; CeA, central nucleus of the amygdala; CoA, cortical nucleus of the amygdala; MeA, medial nucleus of the amygdala. (b) Representation of photomicrograph processing for morphology survey. Cells are counted from original images by blinded scorers. Images are then deconvoluted in order to visualize only DAB positive area, then thresholded to binary images at a predetermined intensity to measure Iba1 proportional area. Deconvoluted images are filtered and again made binary, in order to skeletonize cellular processes. The Analyze Skeleton plugin is then used to identify and measure number of branches and branch length.