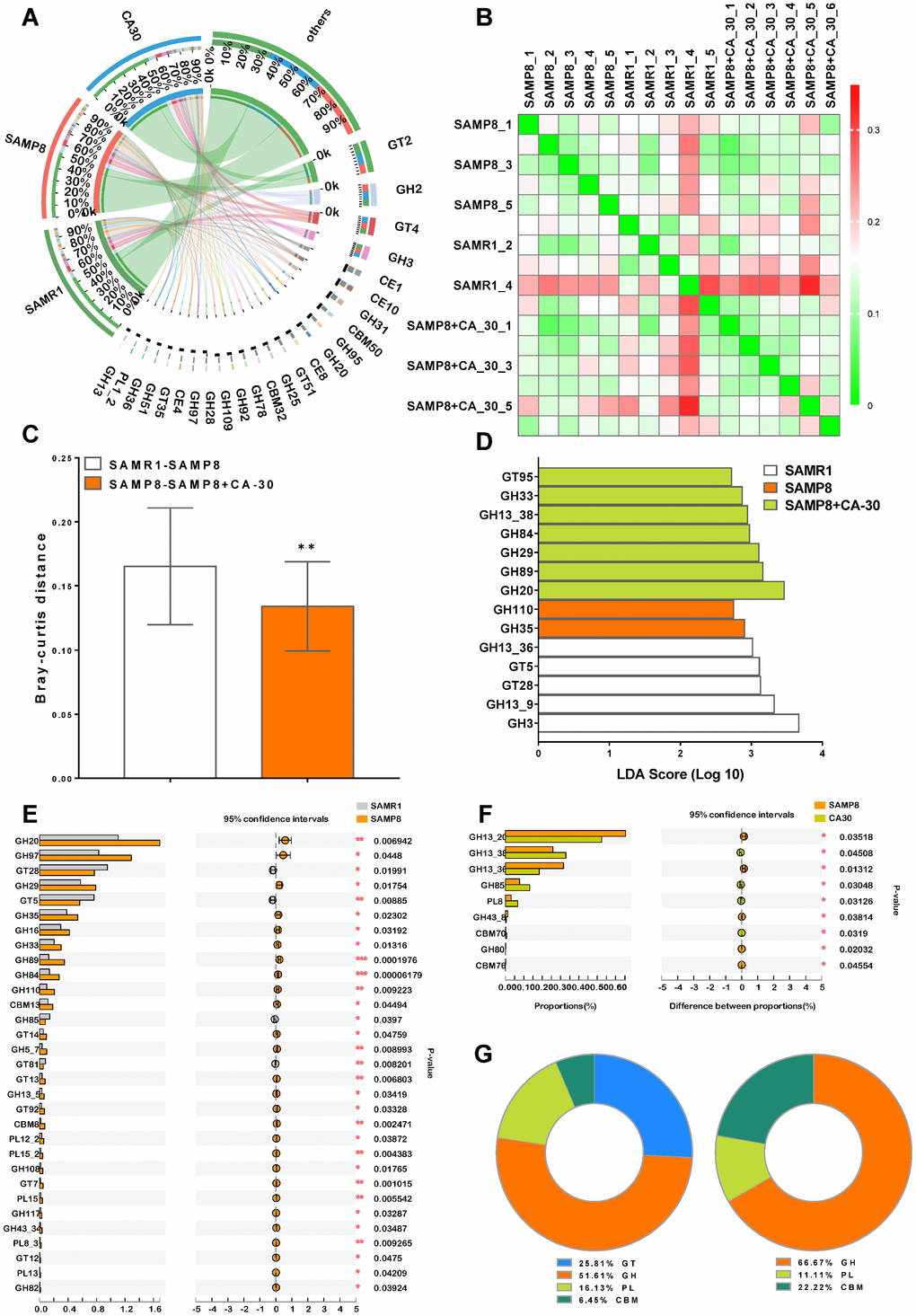
Figure 7.CA-30 modulated the genetic capacity for carbohydrate utilization of gut microbiota. Circos plot of the glycosyltransferases (GT), glycoside hydrolase (GH), polysaccharide lyase (PL), carbohydrate-binding module (CBM), and carbohydrate esterase (CE) families of carbohydrate degrading enzymes found in the metagenome using the carbohydrate-active enzyme (CAZy) database and percent of community abundance of the CAZy family level (A). Bray-Curtis distance heat map at the CAZy family level with the highest frequency and relative abundance, as calculated via the unweighted pair group method with arithmetic means (B) and Bray-Curtis distances among the mice (C). **P<0.01, versus the SAMR1-SAMP8 mice by unpaired Student's t-tests. The most differentially abundant CAZy family in each group was identified by linear discriminant analysis scores generated via linear discriminant analysis effect size analyses (D). Comparison of the relative abundances of the dominant CAZy Family in all groups (E and F). *P<0.05, **P<0.01, ***P<0.01, versus the SAMR1 or SAMP8 mice by unpaired Student's t-tests. The ratio of significantly changed CAZy classes between the SAMR1 and SAMP8 groups and the SAMP8 and CA-30 groups (G). All values are means ± S.D. n=5-6.