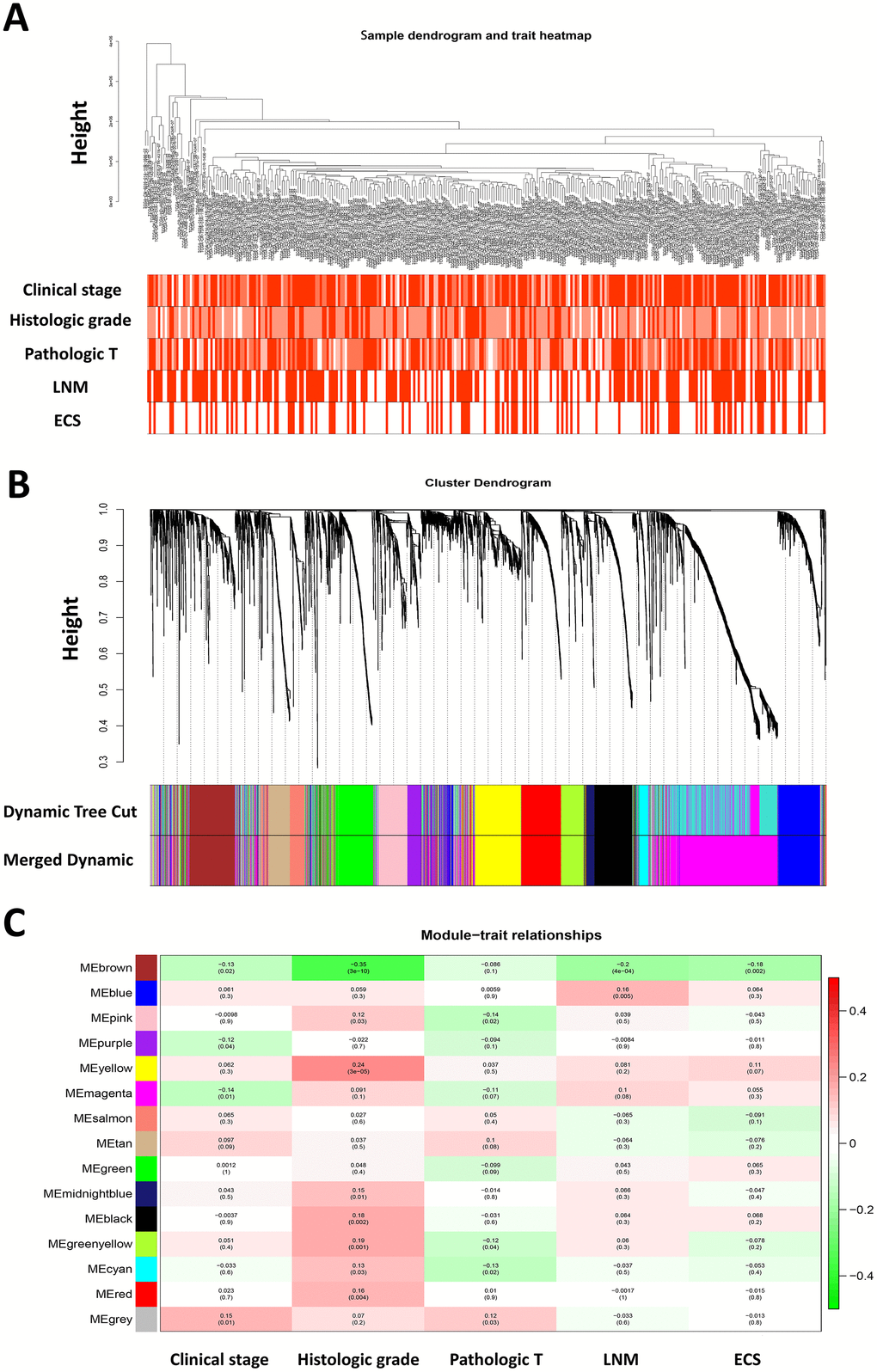
Figure 1.Weighted gene co-expression network analysis and identification of modules associated with the progression of HNSCC. (A) Clustering dendrogram of 299 HNSCC samples and the clinical traits. The clustering was based on the expression data of differentially expressed genes between tumor samples and normal samples in HNSCC. The color intensity was proportional to more advanced clinical stage as well as higher histologic grade and pathologic T stage. In lymph node metastasis (LNM) and nodal extracapsular spread (ECS), the red color represented pathologic nodal metastasis and nodal capsular spread. (B) Dendrogram of all differentially expressed genes clustered based on a dissimilarity measure (1-TOM). (C) Heatmap of the correlation between module eigengenes and disease progression features of HNSCC. The upper number in each cell refers to the correlation coefficient of each module in the trait, and the lower number is the corresponding p-value.