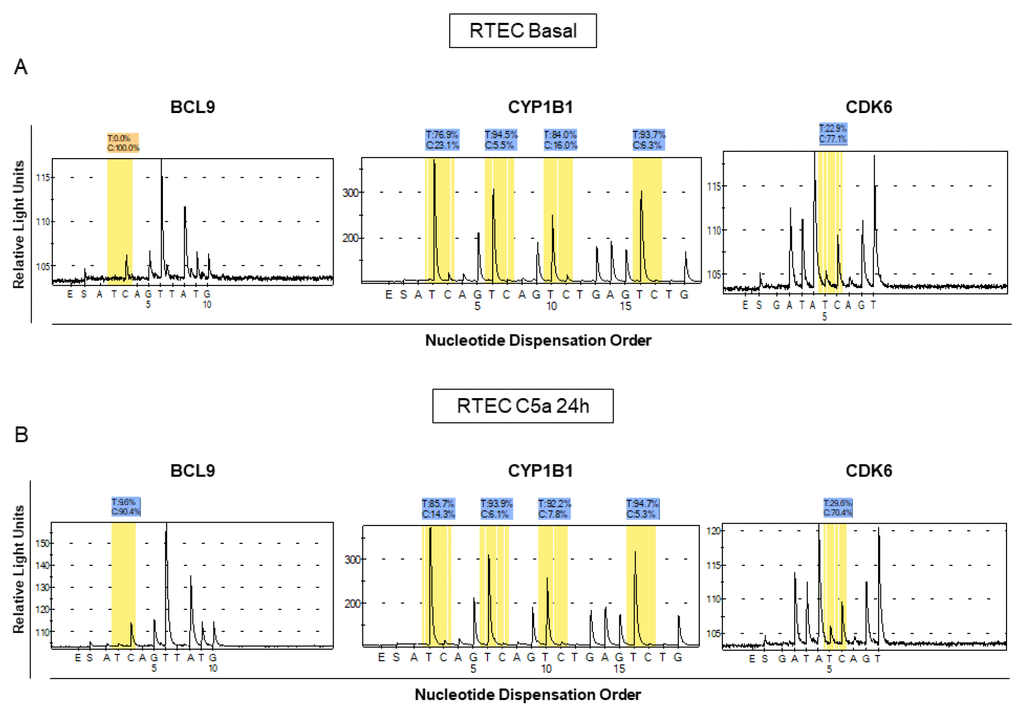
Figure 2.Analysis of methylation levels for BCL9, CYP1B1 and CDK6 in different lots of C5a-stimulated RTEC compared to the respective basal conditions. Pyrosequencing assays were performed on the same regions that we found to be methylated in the whole-genome assay. The pyrosequencing assays confirmed that the DNA of these regions were differentially methylated in C5a-stimulated RTEC (B panel) compared to basal condition (A panel). Representative pyrograms show methylation levels of the indicated CpG sites in the gene promoter.