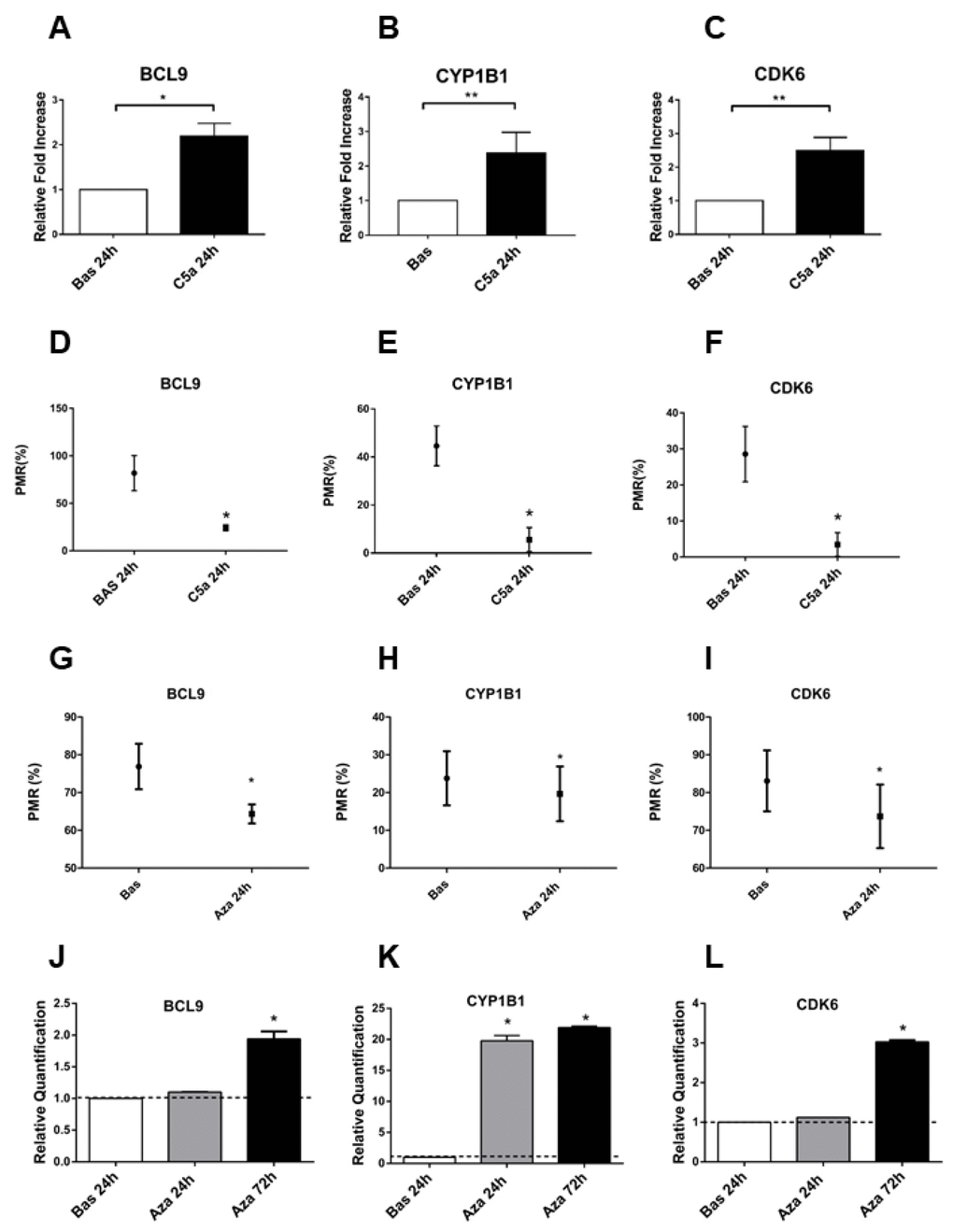
Figure 3.BCL9, CYP1B1 and CDK6 gene expression is regulated by the DNA methylation. (A–C) Gene expression of BCL9, CYP1B1 and CDK6 evaluated by qRT-PCR in the C5a stimulated-RTEC and compared to normal RTEC cultured for 24h. The gene expression validated the genes analyzed for the methylation using the pyrosequencing assay. Expression levels were significantly different in C5a-stimulated RTEC compared with normal cells. Gene expression levels were normalized to the housekeeping gene GAPDH. (D–F) BCL9, CYP1B1 and CDK6 methylation levels of C5a-stimulated RTEC compared to basal condition. Results are means±SD, n=3. (G–I) Gene expression of BCL9, CYP1B1 and CDK6 of C5a stimulated-RTEC not treated or treated with 1 μM 5-aza-2’-deoxycytidine. The DNA demethylation agent reduced the methylation levels in the promoters of the three genes. The DNA methylation status of these three DNA regions was determined by qMSP real-time analysis. The degree of fully methylated molecules at a specific locus was expressed as a PMR index. The percentage PMR was calculated as described in the Materials and methods. Qiagen methylation control DNA was used as full methylated reference. (J–L) Gene expression of BCL9, CYP1B1 and CDK6 of C5a stimulated-RTEC not treated or treated with 1 μM aza. Results are means±SD, n = 3.*p<0.05.