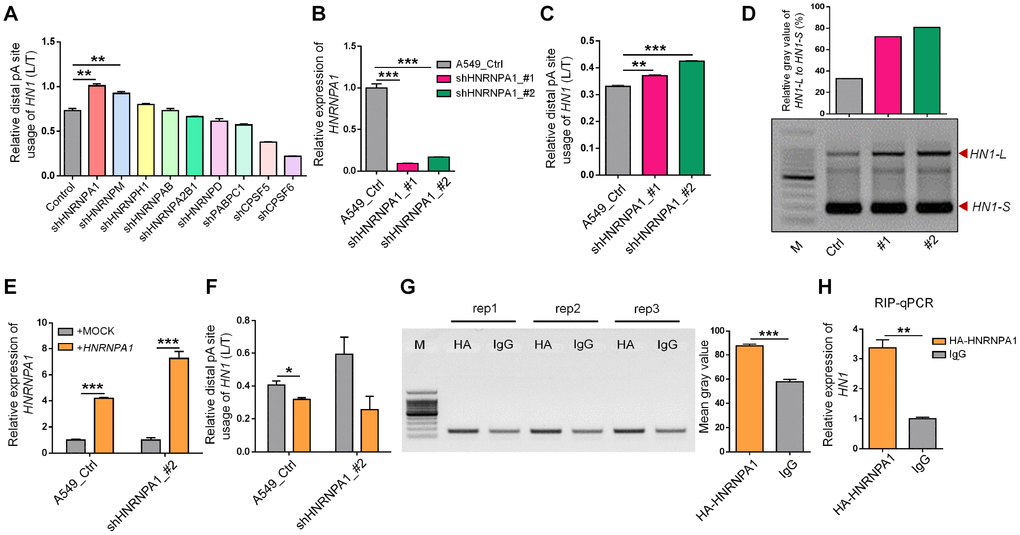
Figure 4.HNRNPA1 binds to mRNA of HN1 and regulates its 3′ UTR length changes. (A) The relative HN1-L expression to total mRNA levels (L/T) evaluated by qRT-PCR upon knockdown of various genes encoding candidate RNA binding proteins in HEK293T. ** represents p < 0.01 based on t-test with three qPCR reactions. (B) Knockdown of HNRNPA1 by two shRNAs (shHNRNPA1_#1, shHNRNPA1_#2) was confirmed by qRT-PCR in A549 cells. *** represents p < 0.001 based on t-test with three qPCR reactions. (C) Higher usage of HN1-L to total HN1 mRNA (L/T) upon HNRNPA1 knockdown in A549 cells was evaluated by qRT-PCR analysis. ** and *** represent p < 0.01 and p < 0.001, respectively, based on t-test with three qPCR reactions. (D) 3′ RACE assay showed higher abundance of HN1-L compared to HN1-S upon knockdown of HNRNPA1. Relative mean gray value of HN1-L compared with HN1-S was shown at the upper panel by ImageJ. (E-F) HNRNPA1 overexpression reversed HNRNPA1-KD induced 3′ UTR lengthening of HN1. Overexpression of HNRNPA1 was confirmed in both HNRNPA1-KD and control A549 cells (E). Decreased usage of HN1-L compared to total HN1 mRNA (L/T) was detected by qRT-PCR upon overexpression of HNRNPA1 in A549 cell (F). * and *** represent p < 0.05 and p < 0.001, respectively, based on t-test with three qPCR reactions. (G) RNA binding protein immunoprecipitation coupled with PCR (RIP-PCR) was performed in A549 cells transfected with the HA-tagged HNRNPA1-overexpression plasmid. Immunoprecipitated RNA with either HA-antibody or control IgG was reversely transcribed and amplified with primer pairs specific for HN1 mRNA. Agarose gel of three independent RIP-PCR experiments (rep1, 2, 3) was shown at left and mean gray value was shown at right. *** represents p < 0.001 based on t-test with three RIP-PCR reactions at the left. (H) Quantitative PCR for immunoprecipitated RNA (RIP-qPCR) to test the enrichment of HN1 mRNA in HA-antibody compared to control IgG in A549 cells. ** represents p < 0.01 based on t-test with three qPCR reactions.