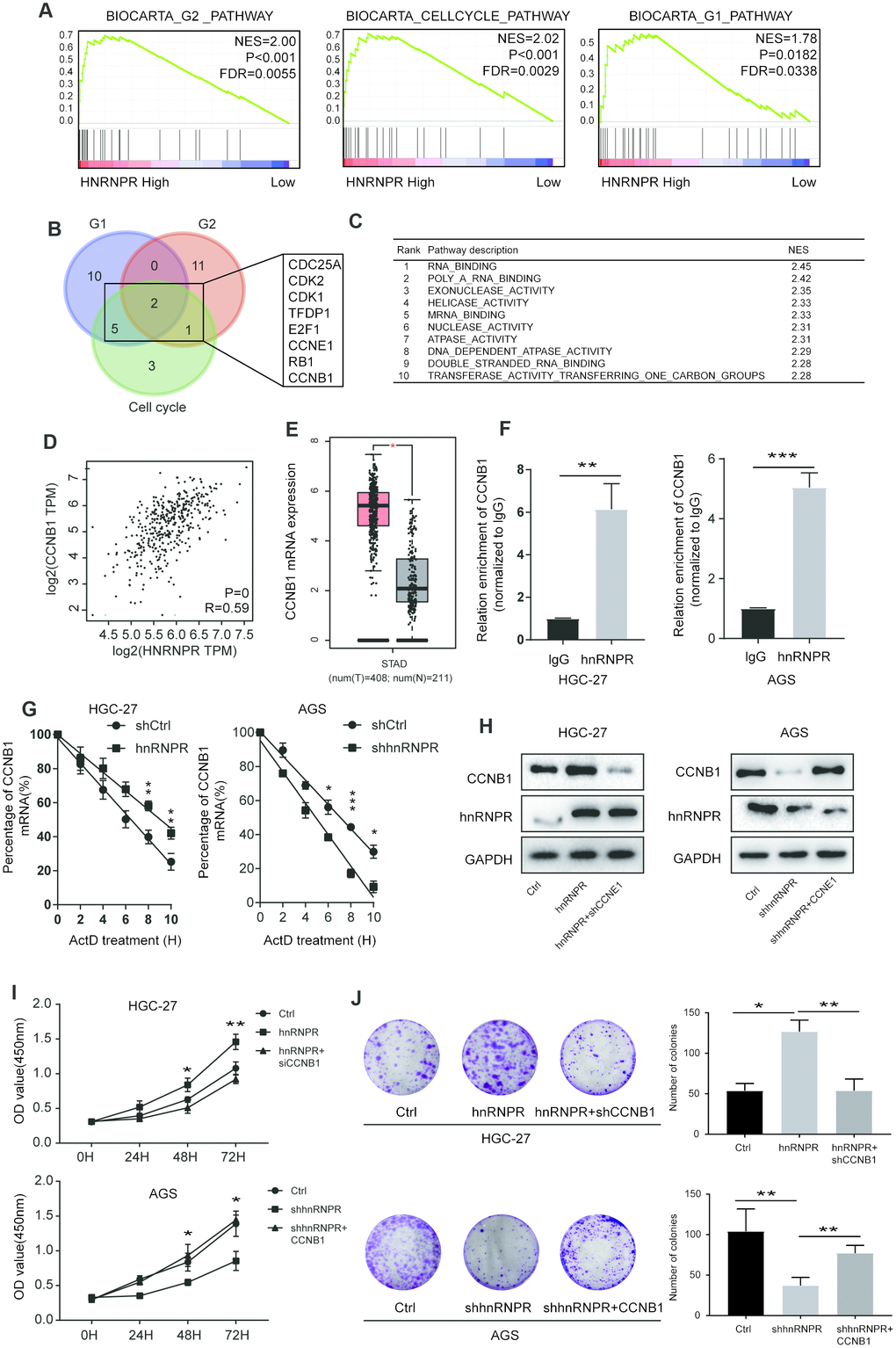
Figure 3.hnRNPR promoted gastric cell proliferation by binding CCNB1 mRNA. (A) GSEA showed that high hnRNPR expression was positively correlated with cell cycle pathway, G2 pathway, and G1 pathway. (B) Venn graph revealed that CDC25A, CDK2, CDK1 TFDP1, E2F1, CCNE1, RB1, CCNB1 overlapped in the three groups. (C) Pathway enrichment analysis showed that “RNA binding” is the top one with significance. (D) The expression of hnRNPR was positively correlated with the expression of CCNB1. (E) The level of CCNB1 was higher in tumors than that in normal tissues. (F) RIP-PCR indicated that CCNB1 mRNA is significantly increased hnRNPR groups in HGC-27 and AGS cell lines. (G) CCNB1 mRNA stability analysis in HGC-27 and AGS cells after actinomycin D (ActD) treatment. Cells were transfected with hnRNPR or shhnRNPR or a control. Cells were harvested at the indicated timepoints. Expression levels were normalized to “0 h” and GAPDH was used as reference gene. (H) HGC-27-control and HGC-27-hnRNPR cells were transfected with shRNA control or shRNA against CCNB1 for western blot, and AGS-control and AGS-shRNPR cells transfected with control or CCNB1 plasmid for western blot. (I) CCK8 and (J) colony formation assays showed that CCNB1 knockdown partially attenuated the enhanced cell proliferation induced by hnRNPR overexpression in HGC-27 cells, while CCNB1 overexpression partially rescued the inhibition of cell growth induced by hnRNPR silencing in AGS cells. Data are from three independent experiments performed in triplicate. P values were calculated with two-tailed unpaired Student’s t test. The expression correlation was determined with Pearson’s correlation analysis. *, P<0.05, **, P<0.01.