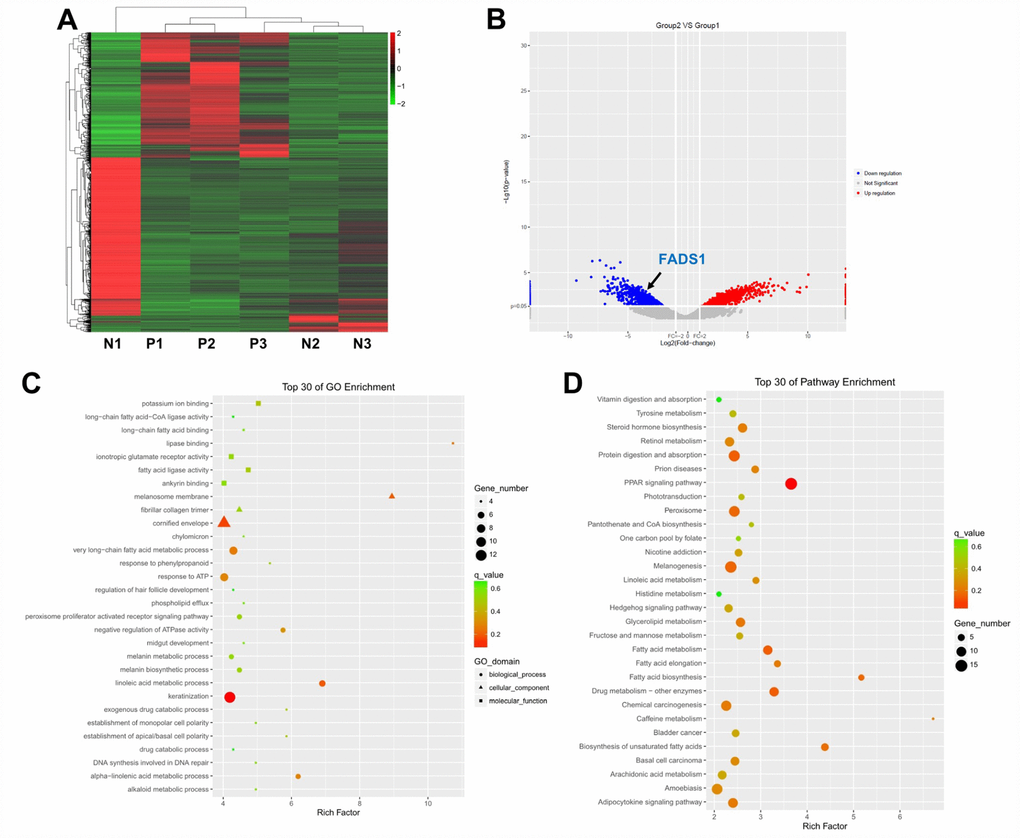
Figure 1.Differential gene expression analysis in clinical vitiligo specimens. (A) Heat map showing gene expression profiles in clinical samples of vitiligo and matched, adjacent normal specimens; each column represents one sample. Red and green indicate upregulation and downregulation, respectively. (B) Distribution of gene transcripts presented as MA plot (log2 fold-change vs log total counts); red points indicate DEGs (FADS1, log2 fold-change = -3.82, -log10 (p-value) = 2.84). (C) Gene ontology (GO) enrichment analysis of DEGs. (D) Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of DEGs. N1, N2, N3: normal specimens; P1, P2, P3: vitiligo samples.