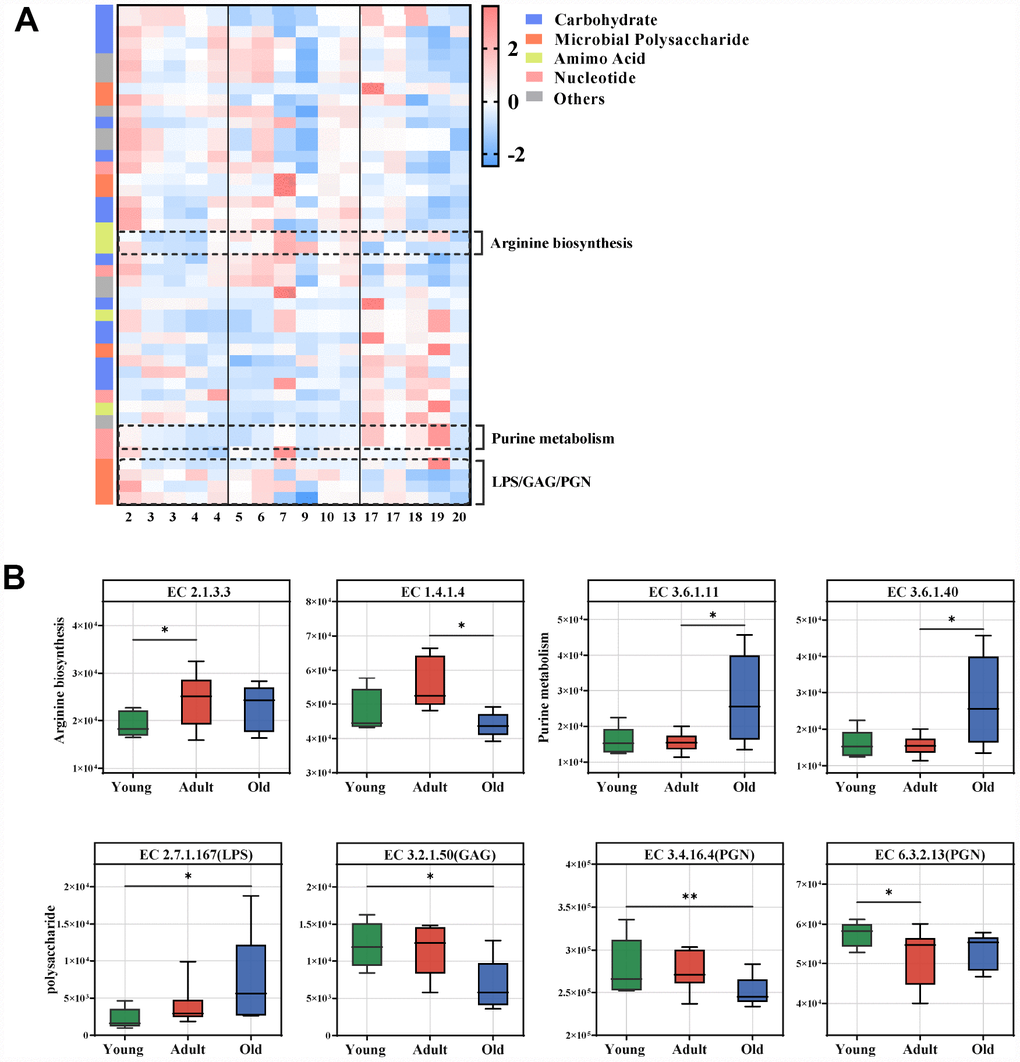
Figure 4.Age-related microbial functions in the KEGG pathway. Different KEGG enzymes identified by LEfSe analysis of the metagenomic sequences (LDA>2.0). (A) Heatmap of the abundances of different enzymes. Enzymes (raw) were sorted by taxa and enriched group, and samples (column) were sorted by age. The intensity of the color (blue to red) indicates the score normalized abundance for each enzyme. (B) Boxplot for the marked KEGG enzymes in different age groups. ECs were classified into pathways for arginine, purine and microbial polysaccharide metabolism. The abundances of different ECs were calculated by reads number. LEfSe was used to detect features with significantly different abundances using the Kruskal–Wallis rank sum test, and LDA was performed to evaluate the effect size of each feature. *P<0.05 **P<0.01. LPS, lipopolysaccharide; GAG, glycosaminoglycan; PGN, peptidoglycan.