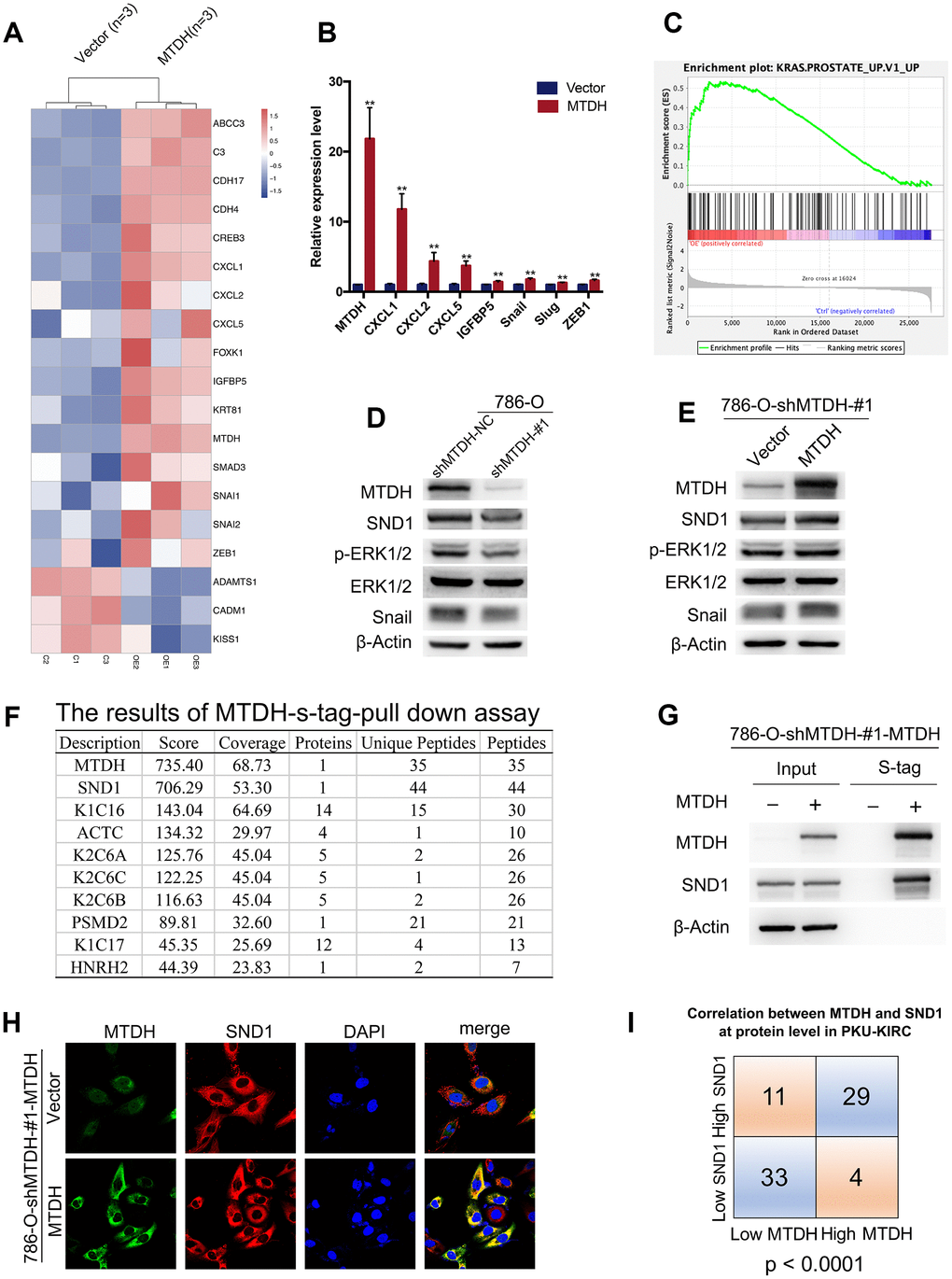
Figure 4.MTDH promotes metastasis by activating ERK signaling and EMT. (A) Heatmap representation of differentially expressed genes identified by RNA-Seq between 786-O-shMTDH-#1-MTDH cells (n = 3) and 786-O-shMTDH-#1-vector Control cells (n = 3). (B) Validation of differentially expressed genes by RT-qPCR. Comparison of mRNA expression of genes in pathways of cancer (CXCL1/2/5 and IGFBP5) and genes in EMT-related pathway (Snail, Slug and ZEB1) between 786-O—shMTDH-#1-MTDH cells and 786-O—shMTDH-#1-vector Control cells. All data are shown as means ± SD. (C) Based on our own RNA sequencing data, genes influenced by MTDH overexpression were mostly enriched in pathways involved with KRAS signaling using Gene Set Enrichment Analysis (GSEA) pathway analysis. (D) Silencing MTDH reduced the protein expression of p-ERK1/2, Snail and SND1 in ccRCC cells. (E) Overexpressing MTDH increased the protein expression of p-ERK1/2, Snail and SND1 in ccRCC cells. (F) The result of mass spectrometry analysis of s-tag pull down assay confirm the interaction between MTDH and SND1 at the protein level. (G) MTDH and SND1 were co-localized, mainly in the cytoplasm. (H) The result of immunoprecipitation revealed that MTDH binds to SND1 at the protein level. (I) The correlation of MTDH and SND1in PKU-KIRC dataset was statistically analyzed (P<0.0001).