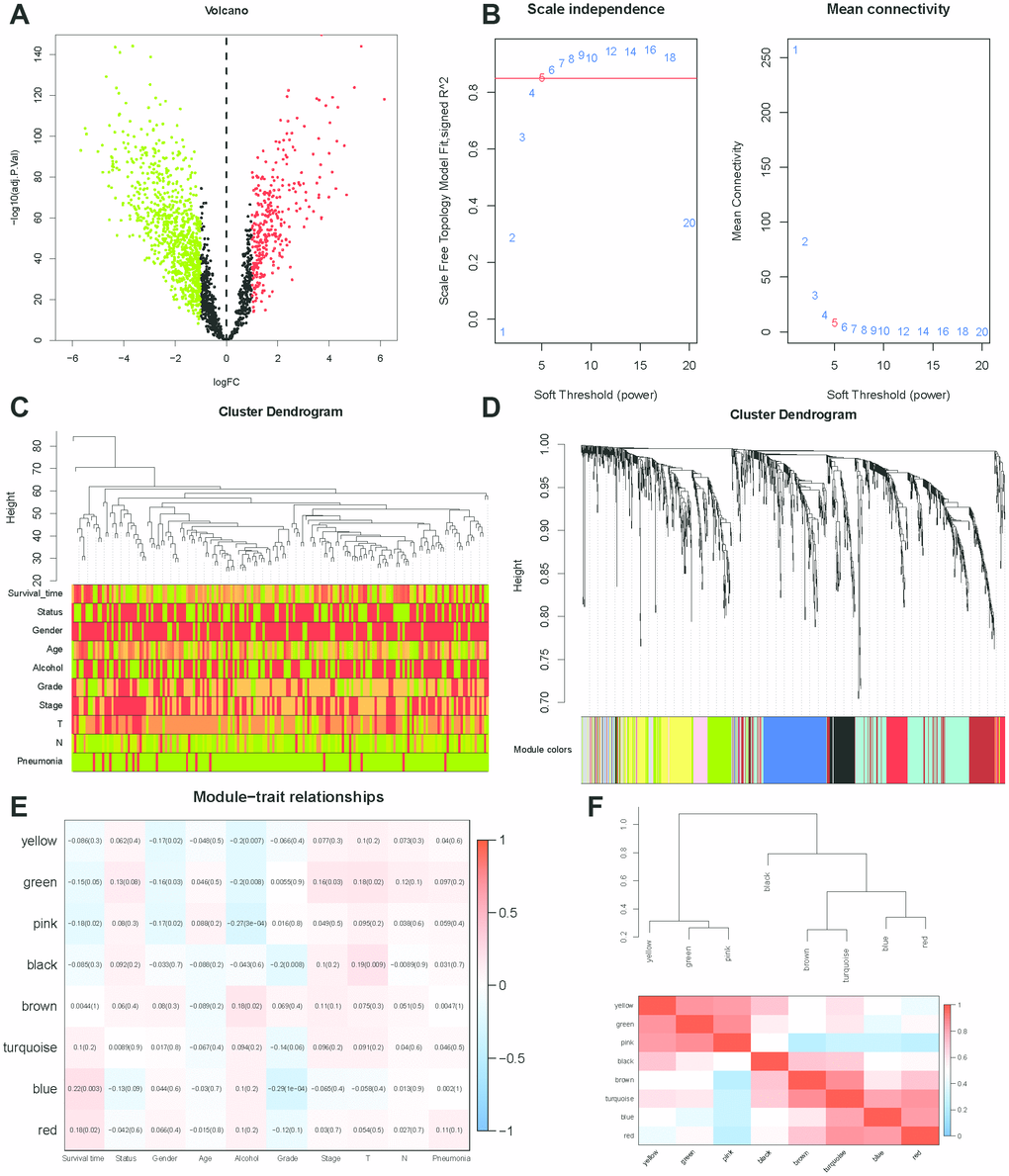
Figure 7.WGCNA analysis. (A) Volcano plot of differentially expressed genes associated with lincRNAs in the signature (correlation coefficient < 0.7, P < 0.001). (B) Analysis of the scale-free topology model fit index for various soft-thresholding powers (β) and the mean connectivity for various soft-thresholding powers. In all, 5 was the most fit power value. (C) Dendrogram of the genes and different clinical factors of ESCC (survival time, status, gender, age, alcohol, grade, stage, T, N, Pneumonia). (D) Dendrogram of the gene modules based on a dissimilarity measure. The branches of the cluster dendrogram correspond to the different gene modules. Each piece of the leaves on the cluster dendrogram corresponds to a gene. (E) Module-trait relationships. Heatmap of the correlation between module eigengenes and clinical characteristics of ESCC. (F) Hierarchical clustering and heatmap of the hub gene network.