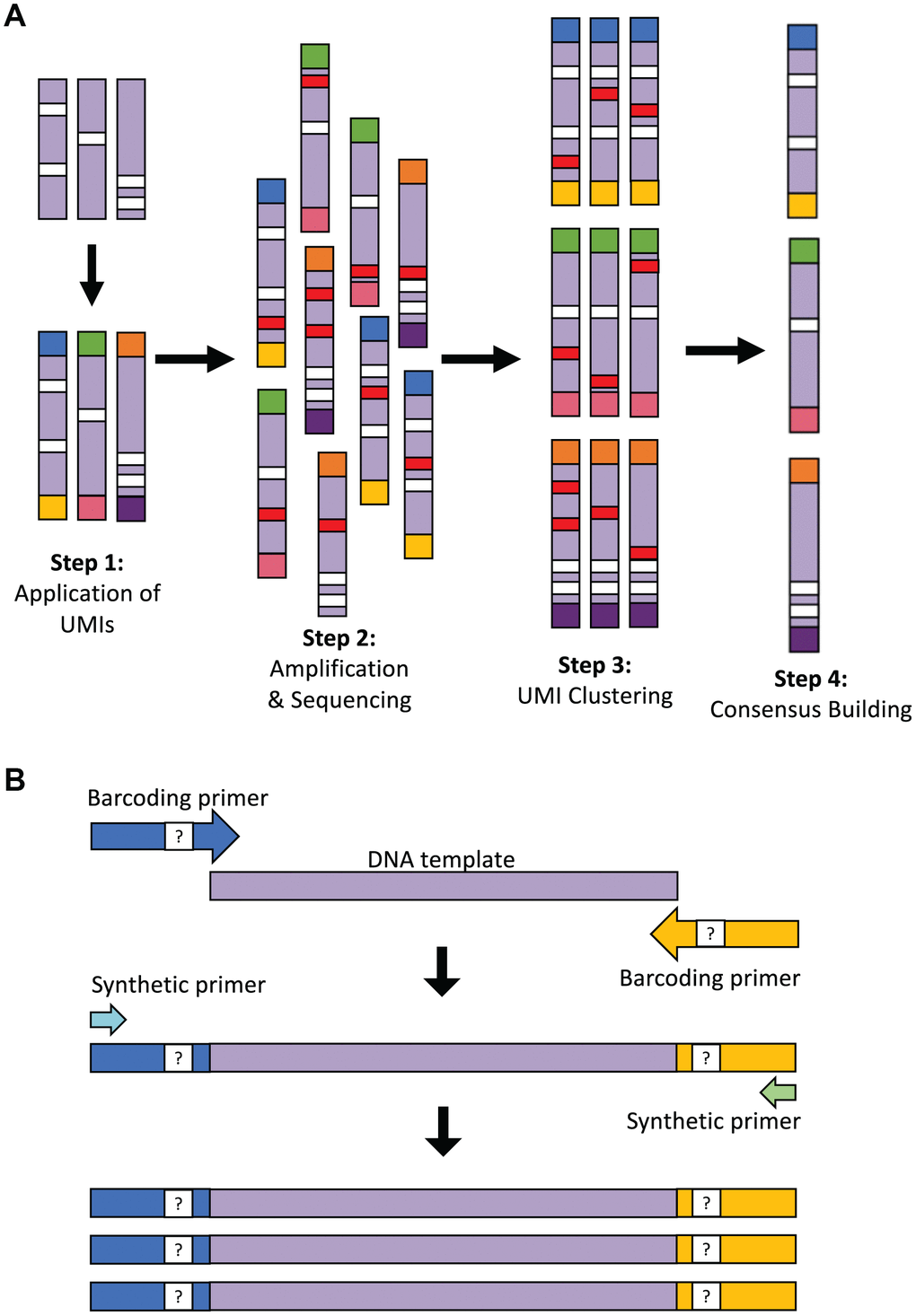
Figure 1.Overview of the LUCS technology. (A) Each individual DNA molecule in a complex mixture, bearing its own unique pattern of mutations (white), has a UMI applied to it via PCR (each UMI represented by a different end-color), which is specific for that molecule (Step 1). The pool of DNA molecules is then amplified and sequenced (Step 2), during which time artefacts (i.e., PCR errors and sequencing errors) are introduced in a random fashion across molecules (red). All reads are then clustered based on their UMI (Step 3), and a consensus read is built for each molecule (Step 4). This final step removes random errors introduced during the process (red) but retains true mutations (white) found in the original molecule and in all amplicons of that molecule. (B) Two-step PCR process for UMI application and dilution. In the first 4 cycles of PCR, the targeted DNA template is amplified by 125-bp oligonucleotide barcoding primers, each containing a random UMI sequence. The initial reaction is then diluted 25-fold within a larger PCR reaction containing only synthetic primers that amplify the UMI-containing molecules after 45 additional cycles of PCR. The resultant elimination of barcoding primer 're-priming' allows for high-resolution pair-end clustering and, in particular, the detection and removal of chimeras (artificial recombinant molecules) caused by PCR jumping.