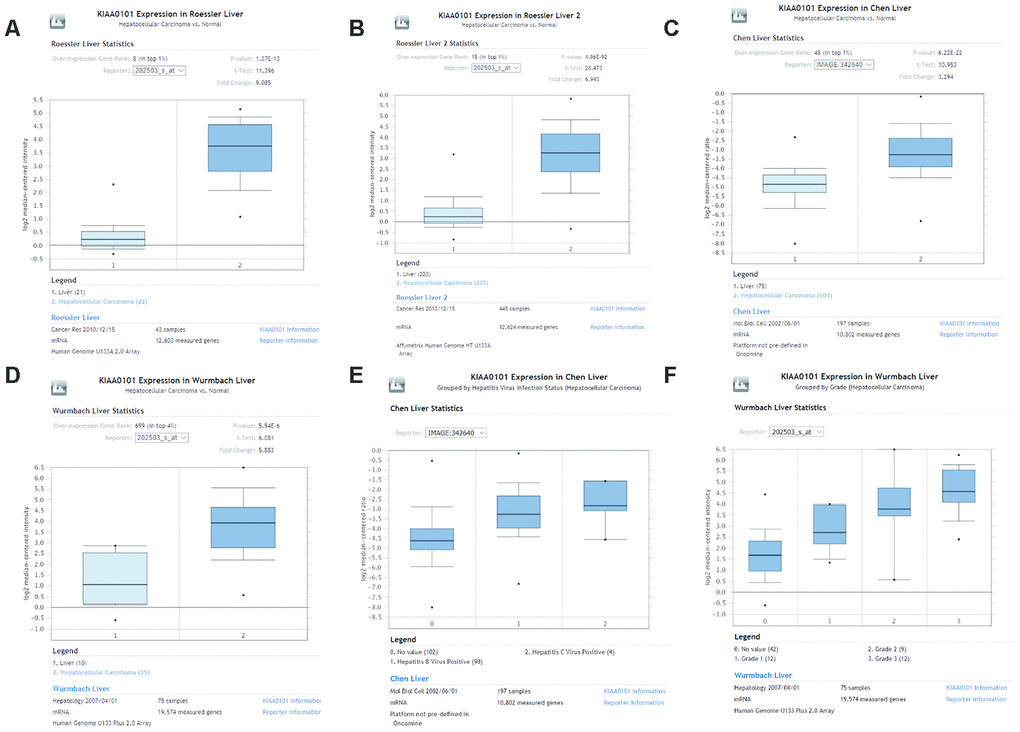
Figure 3.Comparison of KIAA0101 mRNA levels between HCC and normal counterparts by the four unique analyses of ONCOMINE. (A) Comparison of KIAA0101 mRNA levels between noncancer tissues and HCC tumors in Roessler’s study based on Human Genome U133A 2.0 Array in the ONCOMINE database. KIAA0101 presented significantly higher expression in HCC tissues than in noncancer tissues (p=1.37E-13). (B) Comparison of KIAA0101 mRNA levels between noncancer tissues and HCC tumors in Roessler’s study based on the platform of Affymetrix Human Genome HT U133A Array in the ONCOMINE database. KIAA0101 expression was significantly higher in HCC tissues than in noncancer tissues (p=4.96E-92). (C) Comparison of KIAA0101 mRNA levels between noncancer tissues and HCC tumors in Chen’s study in the ONCOMINE database. KIAA0101 presented significantly higher expression in HCC tissues than in noncancer tissues (p= 6.22E-22). (D) Comparison of KIAA0101 mRNA levels between noncancer tissues and HCC tumors in Wurmbach’s study, which was identified in the ONCOMINE database. KIAA0101 presented significantly higher expression in HCC tissues than in noncancer tissues (p= 5.94E-6). (E) KIAA0101 expression in HCC grouped by virus infection status. The HCC samples were no value (102 samples), HBV positive (90 samples), and HCV positive (4 samples) in Chen’s liver dataset. (F) KIAA0101 expression in HCC grouped by tumor grade. The HCC samples are no value (42 samples), grade 1 (12 samples), grade 2 (9 samples), and grade 3 (12 samples) in Wurmbach’s liver dataset. Abbreviations: HCC: hepatocellular carcinoma; HBV: hepatitis B virus; HCV: hepatitis C virus.