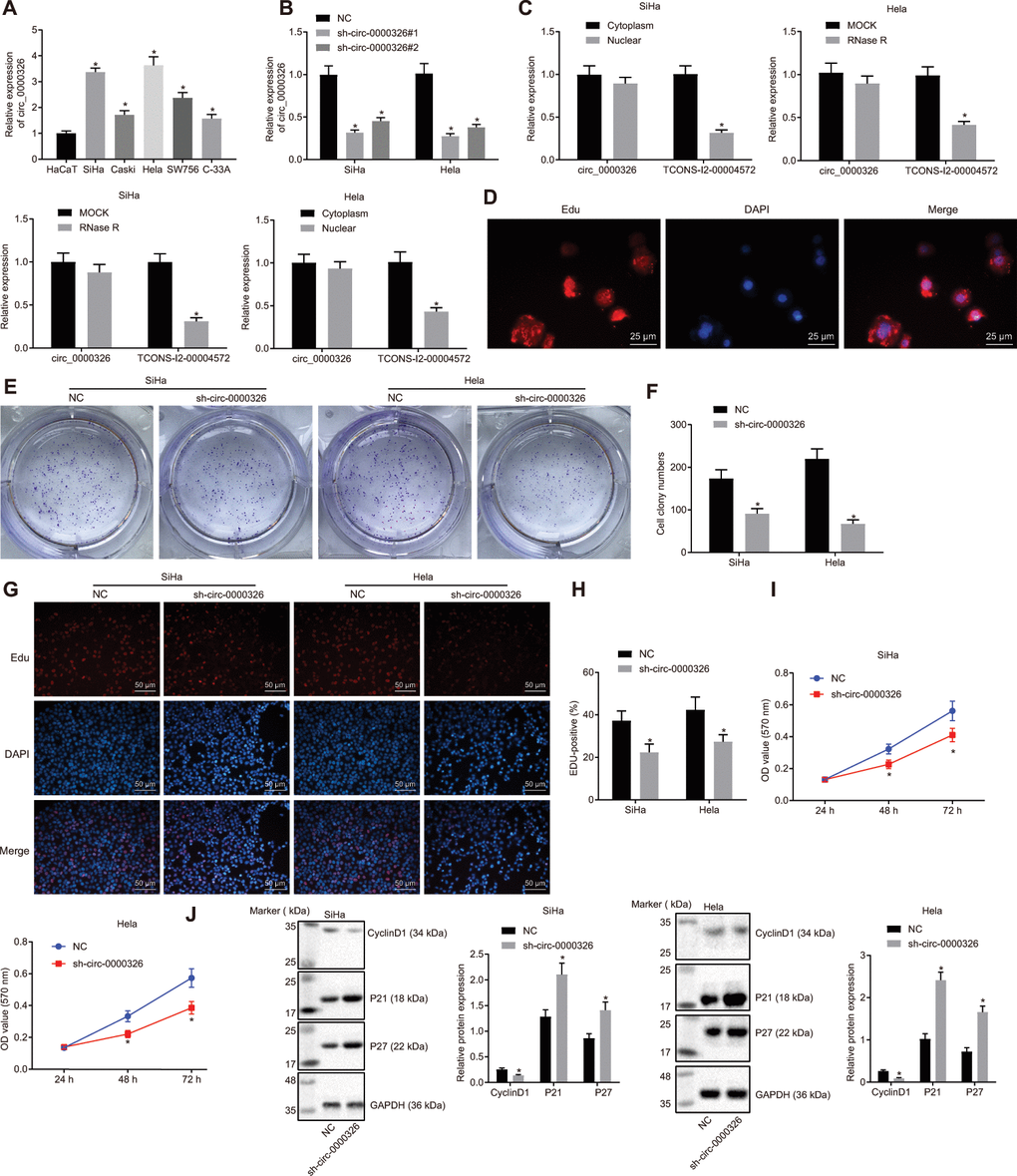
Figure 2.Circ_0000326 promotes proliferation of cervical cancer tissues. (A) Circ_0000326 expression in cervical cancer cell lines upon transfection with shRNA#1, shRNA#2, and shRNA#3. * p < 0.05 vs. the HaCaT cell line or NC. (B) The relative expression of TCONS_l2_00004572 and circ_0000326 after RNase R digestion, * p < 0.05 vs. MOCK. (C) Relative expression of circ_0000326 in nuclear and cytoplasm. * p < 0.05 vs. Cytoplasm. (D) Subcellular localization of circ_0000326 determined by FISH (× 400). (E) Clone formation assay of circ_0000326-silenced cervical cancer cells. (F) The number of cell clones of circ_0000326-silenced cervical cancer cells in clone formation assay. (G) The proliferation SiHa and Hela cell after silence of circ_0000326 determined by EdU assay (× 200). (H) EDU-positive cell rate of SiHa and Hela cells. (I) The cell viability of circ_0000326 silenced SiHa and Hela cells detected by CCK-8. (J) Protein expression of cell cycle-associated proteins (cyclinD1, P21 and P27) in cells after treatment of circ_0000326 determined by Western blot analysis. From figure (D–J), * p < 0.05 vs. NC treatment. Data were expressed as mean ± standard deviation. The data between two groups were analyzed by unpaired t-test with independent sample while the data among multiple groups was analyzed by ANOVA followed by Tukey’ s post hoc test. Data at different time points were analyzed by Two-Way ANOVA.