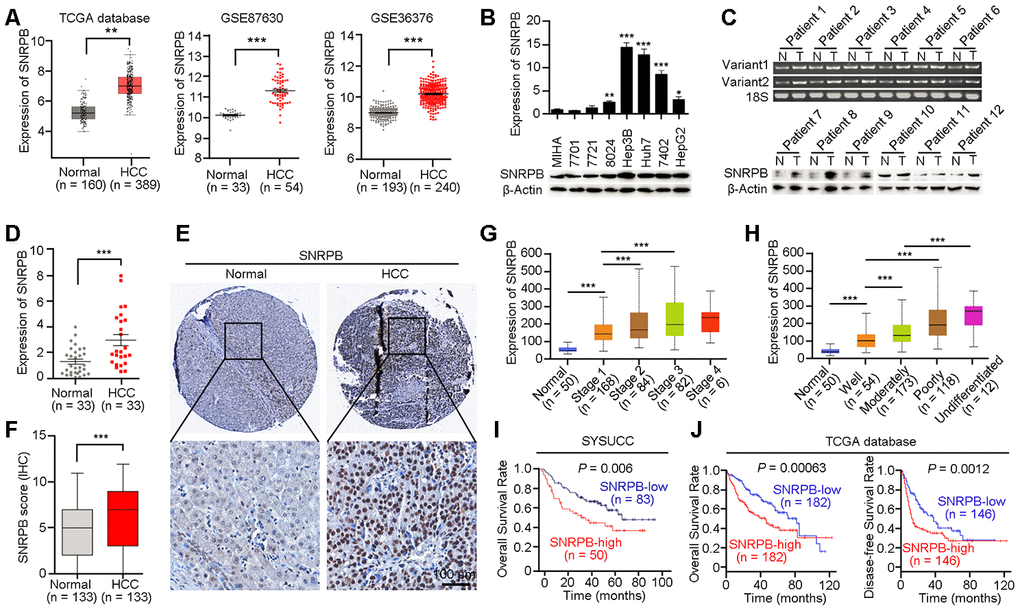
Figure 1.Overexpression of SNRPB predicts poor survival of HCC patients. (A) The expression levels of SNRPB in normal liver tissues and HCC tissues were analyzed based on TCGA database and GEO datasets (GSE87630 and GSE36376). (B) Real-time quantitative PCR (qRT-PCR, upper panel) and western blotting (lower panel) were used to examine the expression of SNRPB in HCC cell lines and immortalized hepatocytes (MIHA). 18S or β-Actin served as the loading controls. (C) The expression levels of SNRPB in paired HCC tumor (T) and normal liver (N) tissues were examined by RT-PCR (upper panel) and western blotting (lower panel). Two splicing variants of SNRPB were analyzed by RT-PCR, and 18S served as the loading control. (D) qRT-PCR was used to examine the expression levels of SNRPB in HCC tissues and the corresponding normal liver tissues (n = 33). (E) Representative images of IHC staining of SNRPB in paired normal liver and HCC tissues. (F) IHC staining scores of SNRPB in HCC tissues and the corresponding normal liver tissues (n = 133). (G, H) SNRPB expression is related to HCC cancer stages (G) and differentiation grades (H) in TCGA database. (I, J) Kaplan–Meier survival curves showed that SNRPB expression level was negatively correlated with HCC prognosis, as analyzed by HCC tissue microarray (I) and TCGA database (J). In all panels, **P < 0.01, ***P < 0.001.