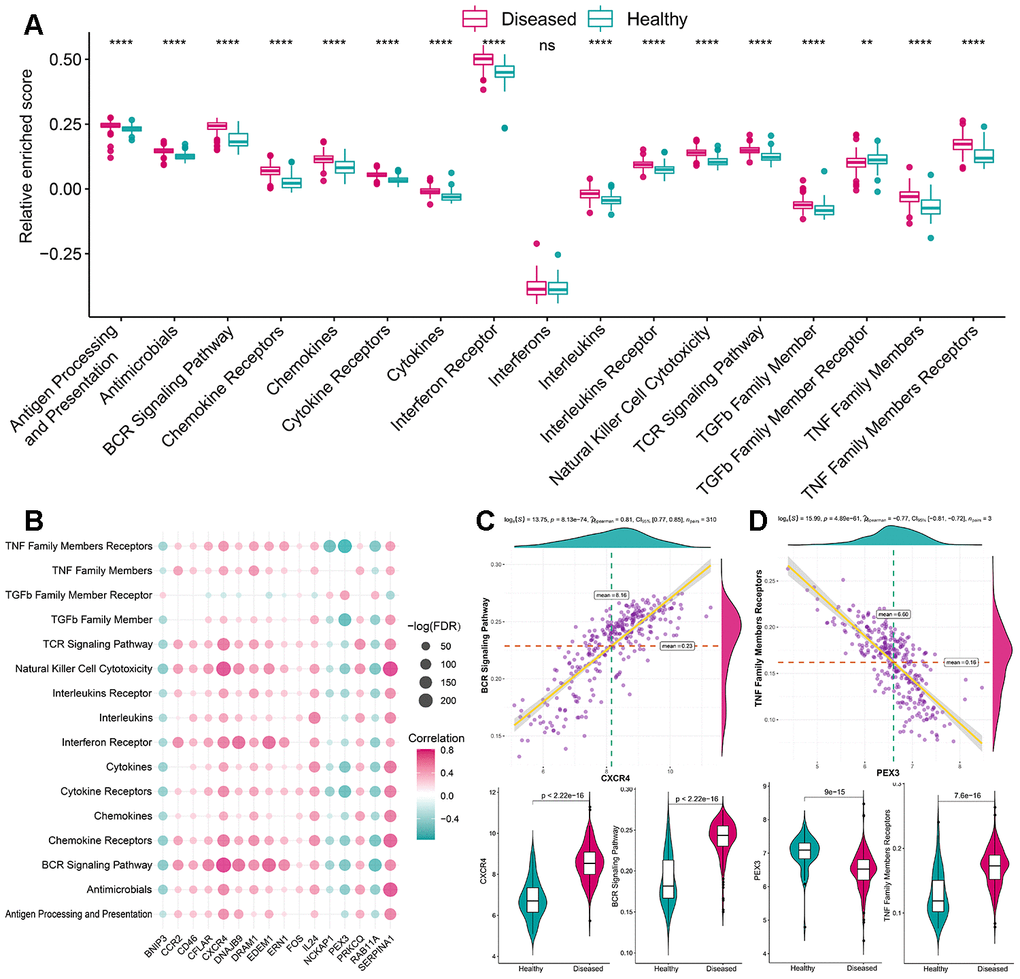
Figure 4.The correlation between immune reaction gene-sets and autophagy genes. (A) The difference in the activity of each immune reaction gene-set between healthy and periodontitis samples. (B) The dot-plot demonstrated the correlations between each dysregulated immune reaction gene-set and each dysregulated autophagy gene. (C) The most positive correlated pair is CXCR4-BCR Signaling Pathway and the expression status or activity status is presented by violin-plot at the left panel, indicating a higher expression of CXCR4 and more active BCR Signaling Pathway reaction were found in periodontitis. (D) The most negatively correlated pair is PEX3-TNF Family Members Receptors and the expression status or activity status is presented by violin-plot at the right panel, indicating a lower expression of PEX3 and more active TNF Family Members Receptors reactions in periodontitis.