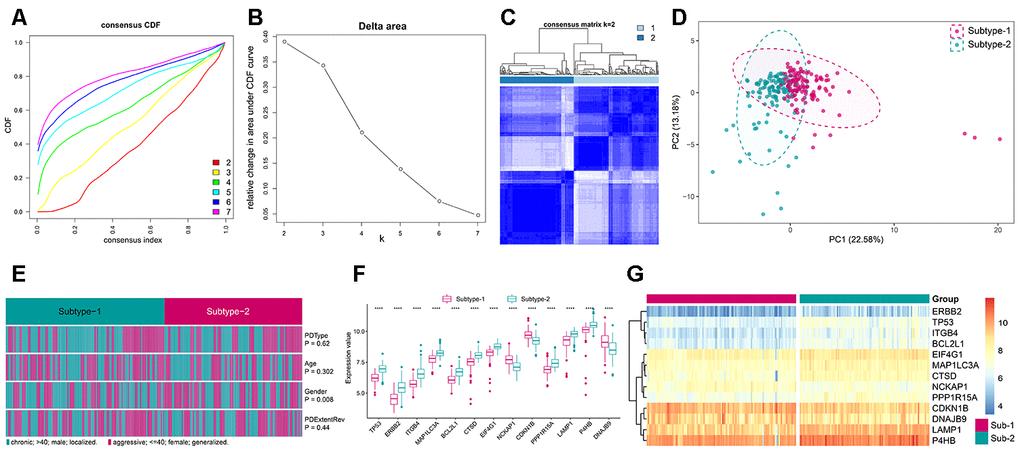
Figure 6.Unsupervised clustering of 208 autophagy genes identifying 2 distinct autophagy-mediated regulation pattern subtypes in periodontitis. (A) Consensus clustering cumulative distribution function (CDF) for k = 2–7. (B) Relative change in area under the CDF curve for k = 2–7. (C) Heatmap of the matrix of co-occurrence proportions for periodontitis samples. (D) Principal component analysis for the transcriptome profiles of 2 autophagy regulation patterns, showing a remarkable difference in transcriptome between different regulation patterns. (E) Comparing of age, gender, periodontitis range and periodontitis type among 2 autophagy regulation patterns. The heatmap illustrates the association of different clinical characters with the 2 subtypes. (F) The expression status of subtype-specific autophagy genes in the two subtypes. (G) Unsupervised clustering of 13 subtype-specific autophagy genes in the 2 regulation patterns.