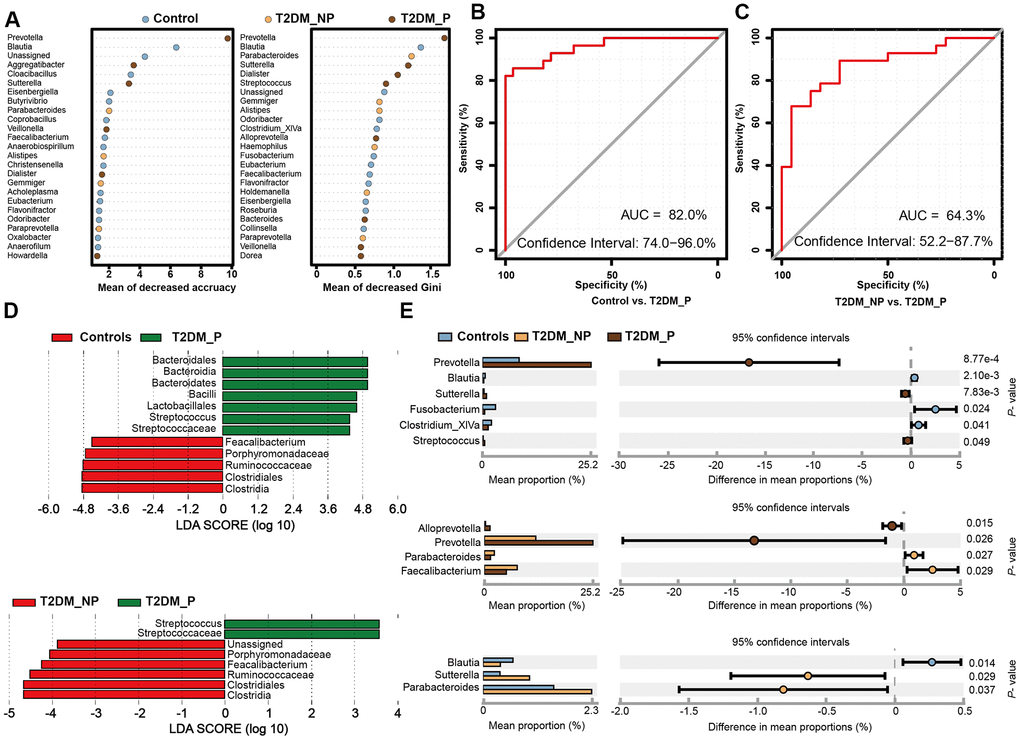
Figure 3.Microbial biomarkers between healthy volunteers and patients with diabetes and periodontitis or diabetes alone. (A) Microbial markers of the T2DM_P group were identified with random forest models. The 34 OTU markers were selected by random forest models as optimal markers to distinguish among groups. (B, C) AUCs were 85.84% (95% CI, 76.3%-95.3%) between T2DM_P and control cohorts (B), and 78.83% (95% CI, 65.4%-92.3%) between T2DM_P and T2DM_NP cohorts (C). (D) Linear discriminant analysis (LDA) revealed differentially abundant taxa as biomarkers using the Kruskal-Wallis test (P < 0.05) with LDA score > 2.0. (E) The Wilcoxon rank-sum test was used for comparisons of relative abundances at the genus level in different groups.