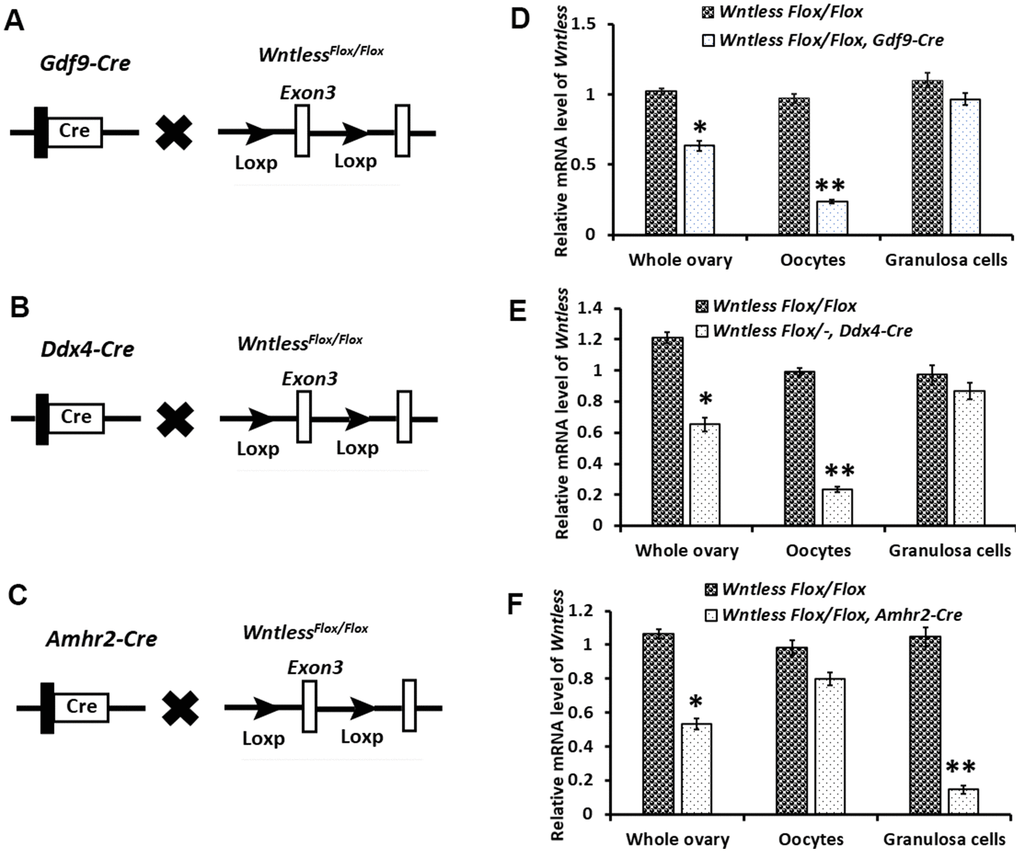
Figure 2.Targeted disruption of the Wntless gene. (A–C) The hybrid scheme used to develop Wntless knockout mice. Mice carrying a targeted Wntless allele (LoxP sites flank Exon 3 of the Wntless allele) were crossed with Gdf9-Cre or Ddx4-Cre or Amhr2-Cre transgenic mice to delete Wntless selectively. The gene knockout was confirmed by PCR genotyping. The isolated genomic DNA from mouse tails was amplified with primer pairs specific for the wildtype (+) (~100 bp) and flox alleles (~200 bp) or different Cre bands (Gdf9-Cre: 326 bp, Ddx4-Cre: 240 bp and Amhr2-Cre: 156 bp). (D–F) qRT-PCR analysis showing the conditional loss of Wntless mRNA in total ovary, oocytes, and granulosa cells extracts of three Wntless knockout mice. Gapdh served as the internal control gene. The data are expressed as the mean ± SEM. *P<0.05, **P<0.01.