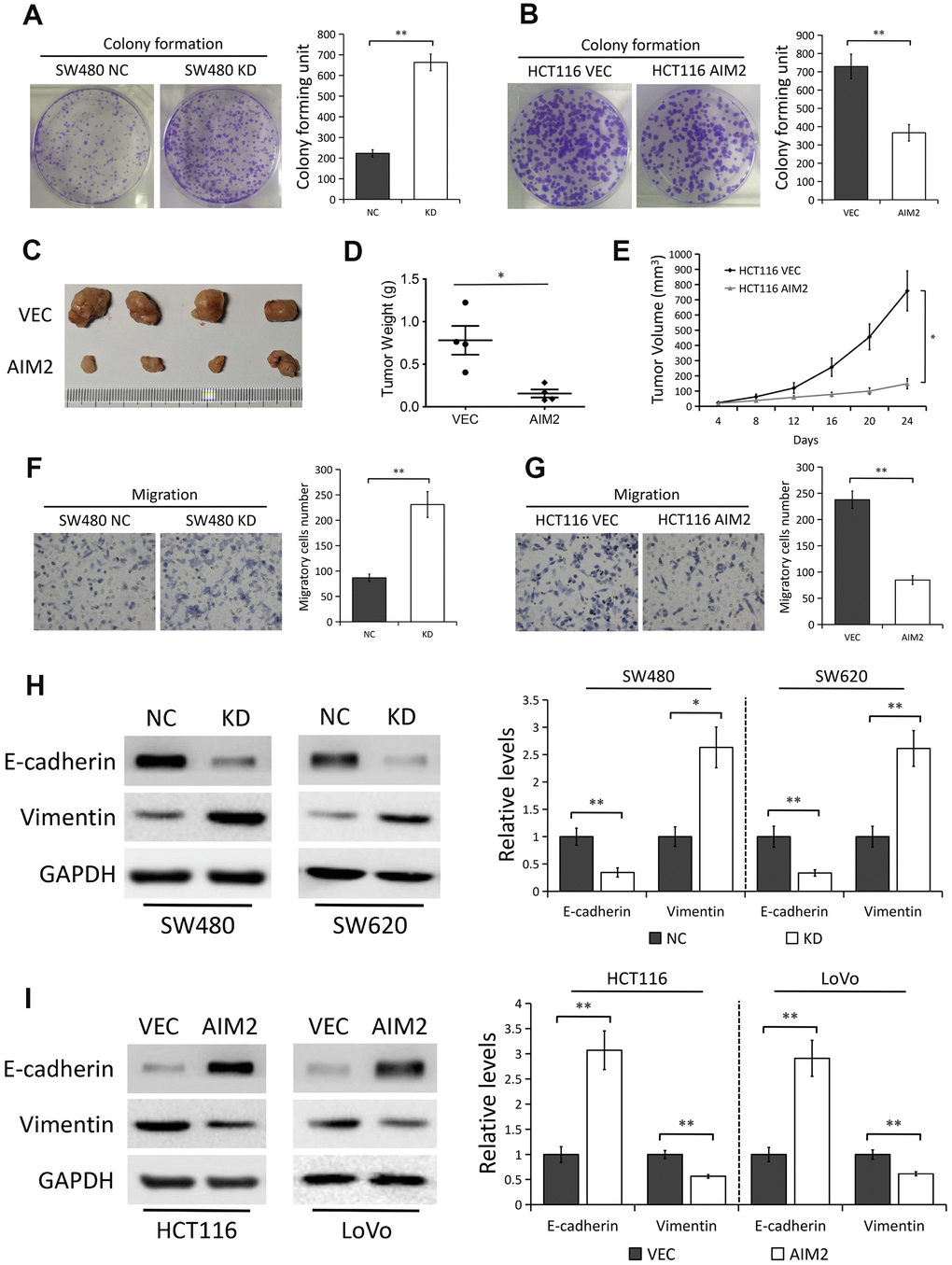
Figure 3.AIM2 plays anti-carcinogenic roles in CRC. (A) Colony formation assays to test viability of SW480 cells stably transfected with control-shRNA (NC) or shRNA against AIM2 (KD). Quantitative analysis results were presented as the mean±SEM (n=3). (B) Colony formation assays to test viability of HCT116 cells stably transfected with empty vector (VEC) or plasmid encoding human AIM2 (AIM2). Quantitative analysis results were presented as the mean±SEM (n=3). (C) Subcutaneous xenograft tumor growth in nude mice (4 per group) was measured and compared in HCT116 (VEC vs. AIM2) cell lines, and the representative image of tumors was shown. (D) Scatter plot analysis of tumor weight of each group was presented. (E) The volumes of the tumors measured every 4 days during the indicated period were shown. (F) Transwell assays to test migration ability of SW480 NC and KD cells. Quantitative analysis results were presented as the mean±SEM (n=3). (G) Transwell assays to test migration ability of HCT116 VEC and AIM2 cells. Quantitative analysis results were presented as the mean±SEM (n=3). (H) Western blots of E-cadherin and Vimentin protein in SW480 and SW620 cells stably transfected with control-shRNA (NC) or shRNA against AIM2 (KD). GAPDH as a loading control. Each experiment was performed at least triplicate and the bands were quantified and presented as the mean±SEM. (I) Western blots of E-cadherin and Vimentin protein in HCT116 and LoVo cells stably transfected with empty vector (VEC) or plasmid encoding human AIM2 (AIM2). GAPDH as a loading control. Each experiment was performed at least triplicate and the bands were quantified and presented as the mean±SEM. *P<0.05, **P<0.01, based on a two-tailed paired Student’s t-test.