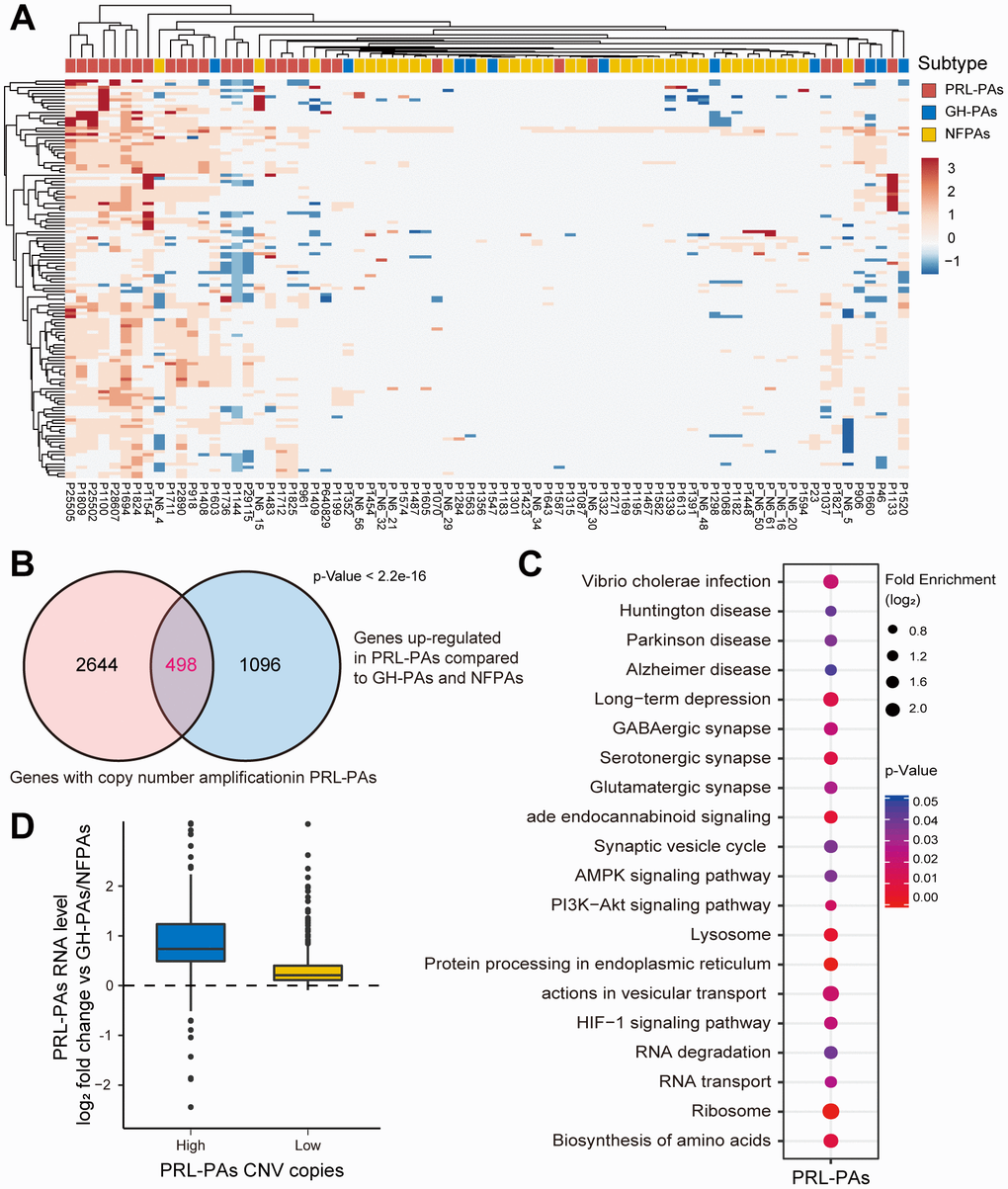
Figure 3.Copy number amplifications in PRL-PAs cause gene transcriptional activation. (A) The heatmap of CNV profiles across PA subtypes. The sample subtype was indicated by color bar above the heatmap, samples were grouped according to the similarity of CNV profiles. The heatmap was colored by CNVs, red indicates gain of copy number and blue indicates loss of copy number. Frequent copy number amplification was observed in PRL-PAs. (B) 498 genes with the copy number amplifications in PRL-PAs overlapped with up-regulated DEGs in the PRL-PAs compared to other subtypes. P value < 2.2e-16 by hypergeometric test. (C) KEGG pathway enrichment analysis of 498 up-regulated genes with both copy number amplifications and transcriptional up-regulation in PRL-PAs, the dot plot shows pathways with a P value < 0.05. (D) PRL-PAs samples were divided into two groups (high CNV and low CNV) according to the clustering results in (A). The boxplot shows the log2 expression fold-changes of PRL-PAs specific up-regulation genes relative to GH-PAs/NFPAs in the high CNV group and low CNV group. The high CNV group exhibited transcriptional up-regulation (Median log2 fold change ~0.75) while the low CNV group did not.