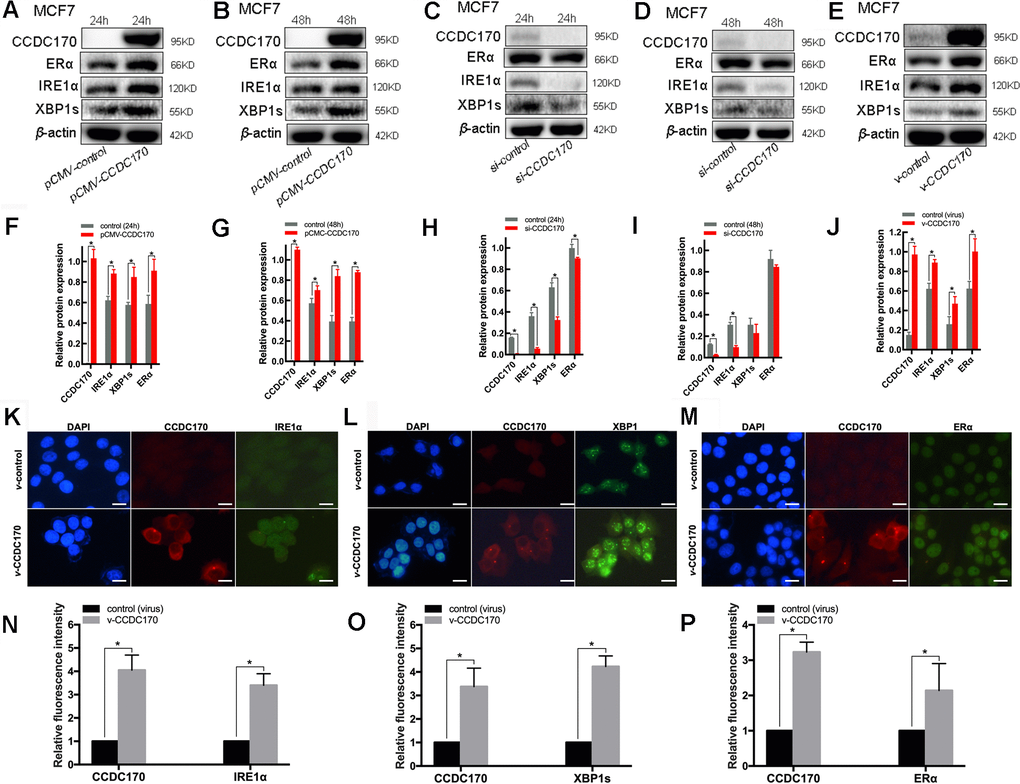
Figure 3.The protein expression of CCDC170, IRE1α and XBP1s in MCF7 breast cancer cells. Representative western blot bands and analysis at 24h (A, F) and 48h (B, G) of CCDC170 up-regulation, 24h (C, H) and 48h (D, I) of CCDC170 down-regulation. Representative western blot bands (E) and analysis (J) in MCF7 breast cancer cells that stably overexpressed CCDC170. β-actin was used as a reference for calculating the relative protein expression. Representative immunofluorescence images and analysis of IRE1α (K, N), XBP1s (L, O) and ERα (M, P) in CCDC170-stably-overexpressing MCF7 cells. Scale bar: 50μm. pCMV-CCDC170(control) represented CCDC170-transiently-overexpressing MCF7 cells and controls. v-CCDC170(control) represented CCDC170-stably-overexpressing MCF7 cells and controls. si-CCDC170(control) represented cells with siRNA-mediated knockdown of CCDC170 and the controls. The error bars presented as mean ± Standard Error of Mean (SEM) with analysis of unpaired Student’s t-test. *P < 0.05, compared with control group.