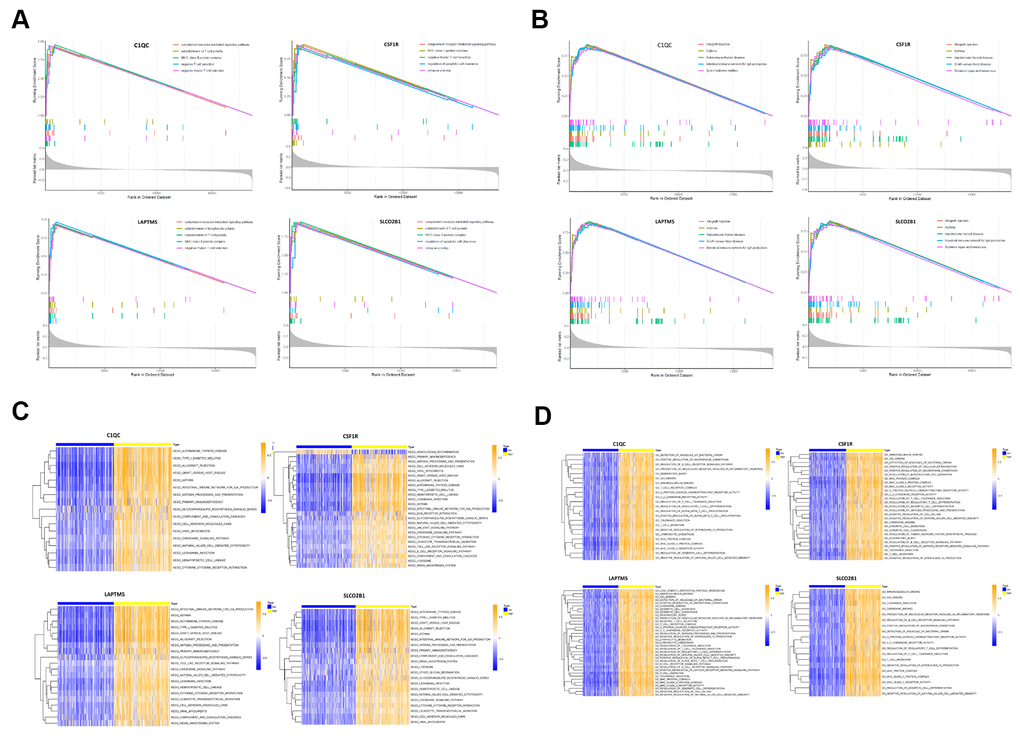
Figure 9.Gene set enrichment analysis (GSEA) and gene set variation analysis (GSVA) of hub genes in the TCGA-LUSC dataset. (A) The plot shows the enriched GO terms based on the GSEA enrichment score in the TCGA-LUSC patients with high expression of each of the four hub genes, LAPTM5, CSF1R, SLCO2B1 and C1QC. (B) The plot shows the enriched KEGG pathways based on the GSEA enrichment score in the TCGA-LUSC patients with high expression of each of the four hub genes, LAPTM5, CSF1R, SLCO2B1 and C1QC. (C) GSVA-derived clustering heatmaps show the enriched GO terms for the LAPTM5, CSF1R, SLCO2B1 and C1QC in the TCGA-LUSC dataset. GO terms with log2 (foldchange) > 0.35 and adjusted P<0.05 are shown. (D) GSVA-derived clustering heatmaps show the enriched KEGG pathways for the LAPTM5, CSF1R, SLCO2B1 and C1QC in the TCGA-LUSC dataset. KEGG signaling pathways with log2 (foldchange) > 0.2 and adjusted P<0.05 are shown.