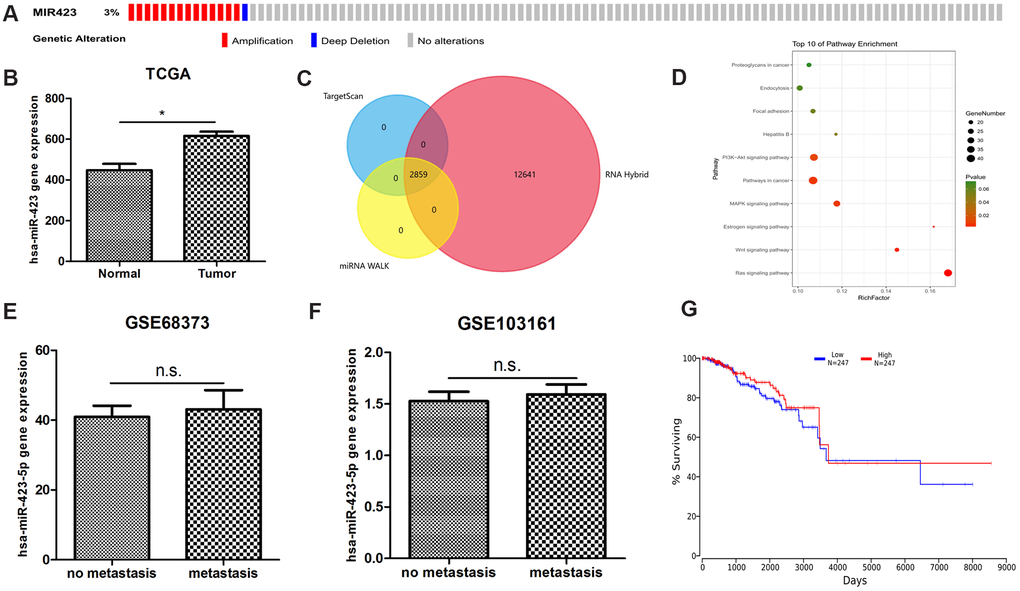
Figure 1.The bioinformatics discovery of breast cancer non-coding RNA biomarkers. (A) The abnormal amplification of hsa-miR-423-5p is shown from the cbioportal analysis. (B) hsa-miR-423-5p is significantly expressed in the TCGA RNA sequencing datasets. (C) A total of 2859 high confidential hsa-miR-423-5p target lncRNAs are found. (D) hsa-miR-423-5p target mRNAs enriched in a number of in cancer-relevant pathways indicate that hsa-miR-423-5p plays a pivotal role in the communication of cancer signaling. (E, F) Two independent microarray data show that hsa-miR-423-5p is not abnormally expressed in breast cancer metastatic tissue compared to the primary tumor, suggesting that hsa-miR-423-5p does not maintain its regulatory role in the advanced stage of breast cancer. (G) Online survival analysis shows that abnormal hsa-miR-423-5p expression is not associated with breast cancer survival.