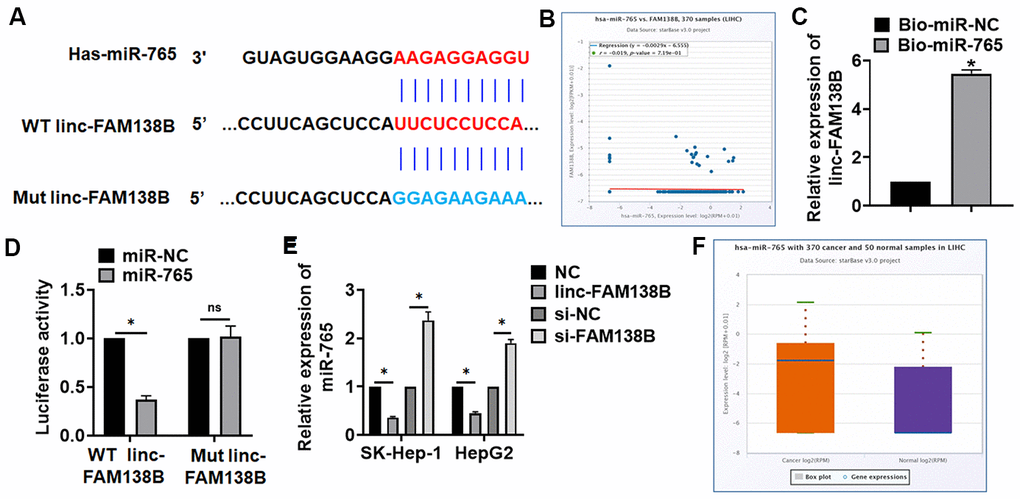
Figure 4.Linc-FAM138B acted as a sponge of miR-765. (A) MiRanda database showing the binding sites of miR-765 with linc-FAM138B, and the mutant sequence of linc-FAM138B. (B) StarBase showed a negative correlation between linc-FAM138B and miR-765 in HCC. (C) Biotinylated miR-765 or NC was transfected into HepG2 cells, and qRT-PCR was performed to detect the enrichment of linc-FAM138B. (D) Wild type and mutant linc-FAM138B was transfected into HEK293 cells with miR-765 or miR-NC, and luciferase assay was to evaluate the binding between miR-765 and linc-FAM138B. (E) SK-HEP-1 and HepG2 cells were transfected with linc-FAM138B plasmid or si-linc-FAM138B or its NC, the mRNA level of miR-765 was detected using qRT-PCR. (F) The expression of miR-765 was identified in HCC patients using database. Data are mean ± SD; *P < 0.05, #P < 0.05 vs cancer exo. Data among multiple groups were analyzed by one-way ANOVA, followed by a Tukey post hoc test. The experiment was repeated in triplicate.