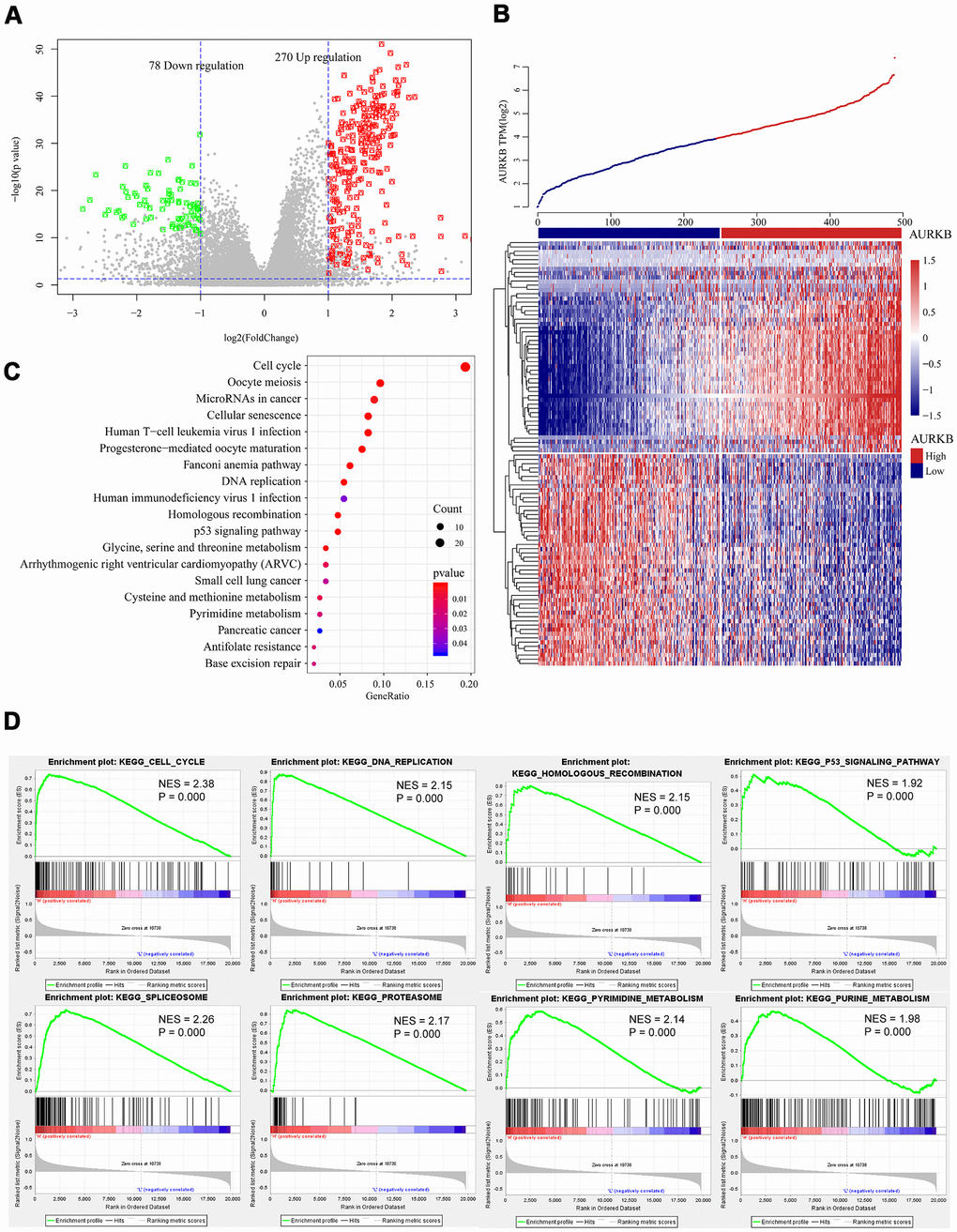
Figure 3.Genome-wide mRNA profiles associated with AURKB expression. (A, B) Volcano plot and heatmap showed the distribution and cluster of DEGs between AURKBhigh and AURKBlow patients. (C) Top 20 pathways were identified through KEGG enrichment analysis by ClusterProfiler R-package, including cell cycle, DNA replication, homologous recombination, and p53 signaling pathway, etc. (D) Significant pathways enriched by GSEA analysis of DEGs between AURKBhigh and AURKBlow patients were shown here: cell cycle, DNA replication, homologous recombination, p53 signaling pathway, spliceosome, proteasome, pyrimidine metabolism, and purine metabolism. Positive values indicate a higher correlation with AURKBhigh patients, while negative values indicate association with AURKBlow patients.