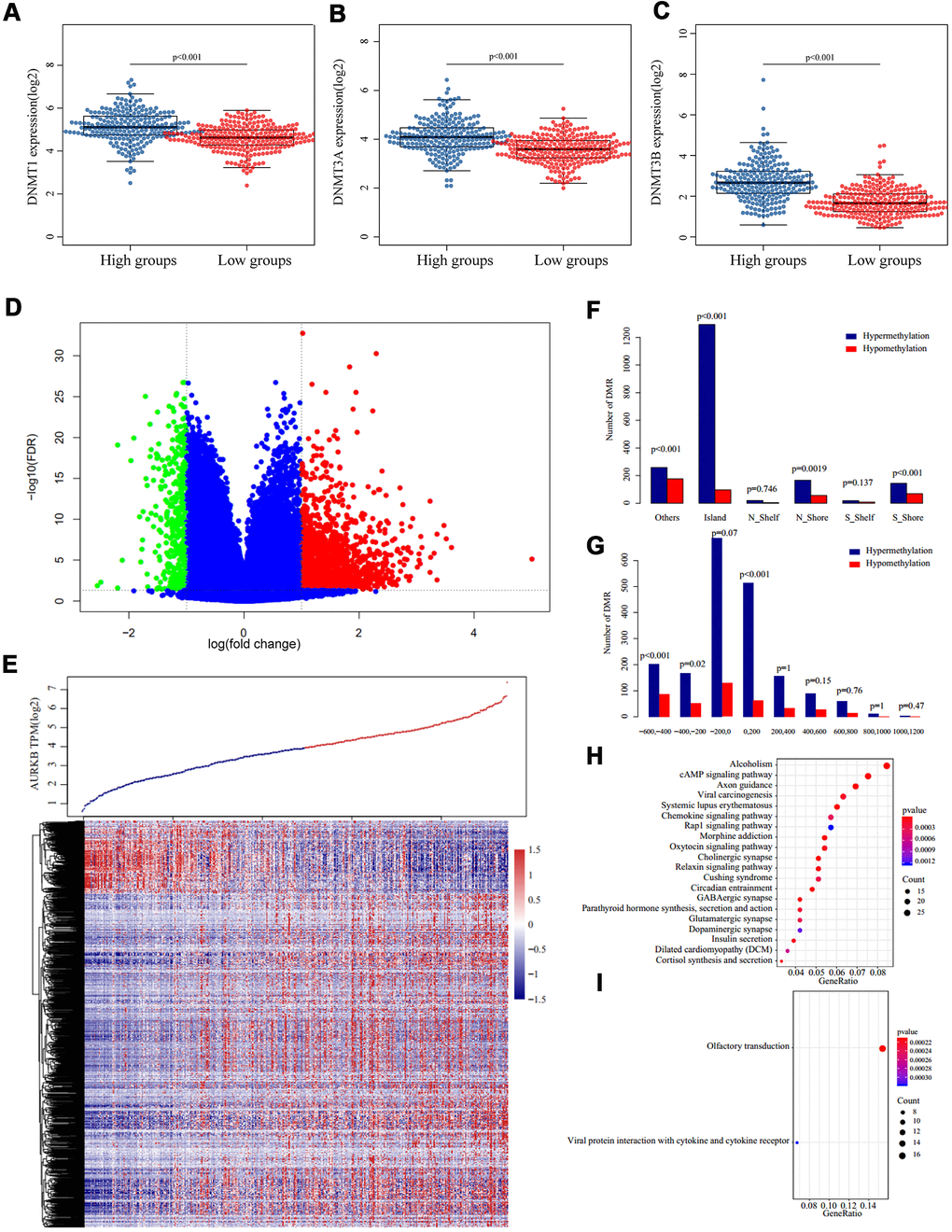
Figure 5.Methylation patterns associated with AURKB expression. (A–C) Expression of three DNA methyltransferases ((A) DNMT1, (B) DNMT3A, (C) DNMT3B) in AURKBhigh and AURKBlow groups, p value as indicated in the figure. (D, E) Volcano plot (D) and heatmap (E) of differentially methylated regions between AURKBhigh and AURKBlow groups (red and green mean significantly DMRs, cutoff fold change 1.3, FDR 0.05). (F, G) Distribution of DMRs on gene’s different structural regions and the distance to TSS. (H, I) KEGG analysis of hypermethylated genes were enriched into 36 pathways, top 20 were listed in (H), while hypomethylated genes were only enriched into two pathways, as shown in (I).