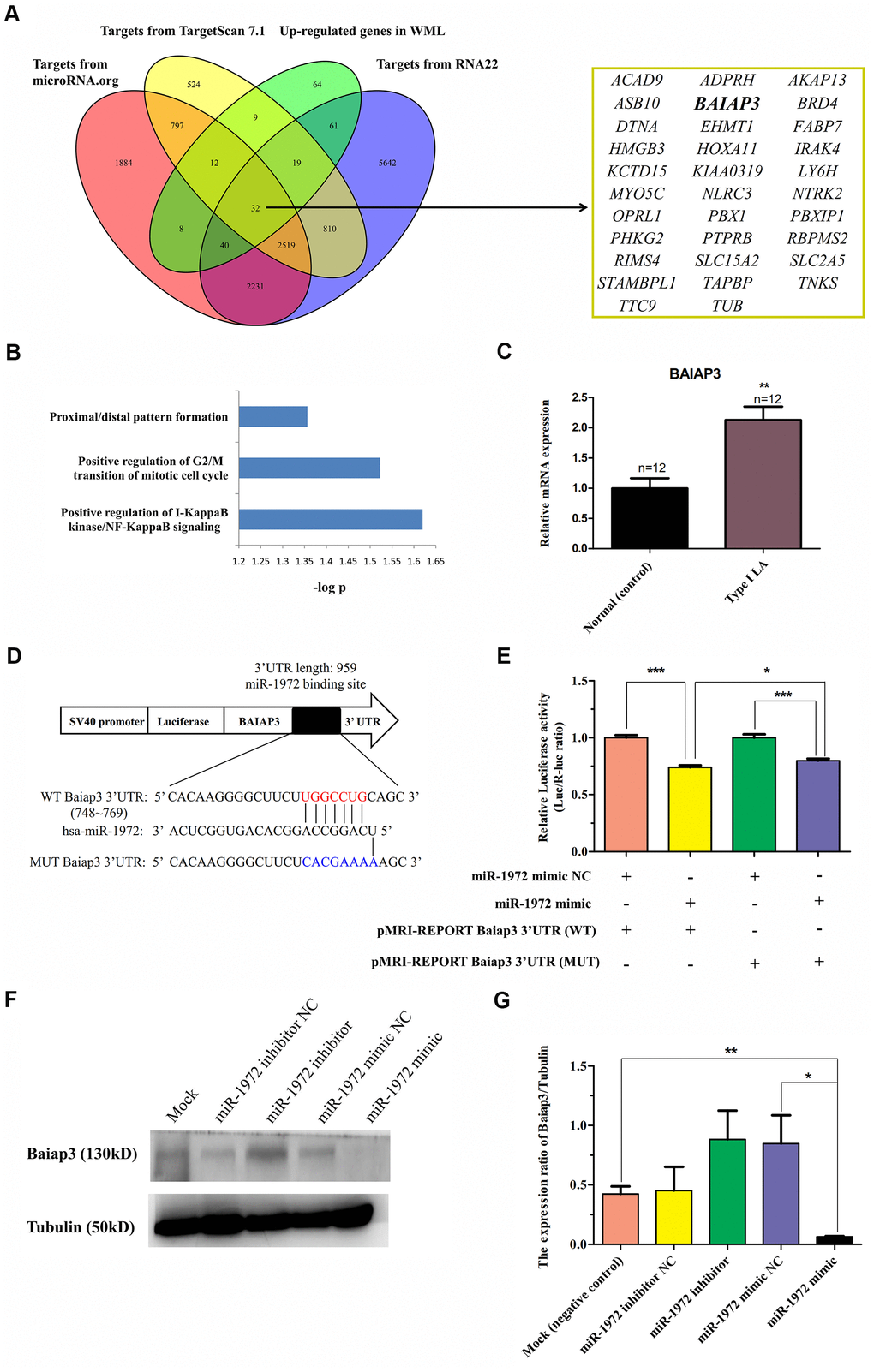
Figure 5.Identification of BAIAP3 as a target of hsa-miR-1972 in LA. (A) The schematic diagram used to search for targets of hsa-miR-1972 in three databases of miRNA targets that were significantly up-regulated in WML tissues. A total of 32 targets of this miRNAs consistently predicted by three predictive tools showed up-regulation in WML tissues compared with non-lesional white matter samples obtained from subjects with WMLs. (B) Functional enrichment analysis of 32 targets of hsa-miR-1972 using DAVID software. Biological processes displaying a statistically significant change with P<0.05 are presented. (C) Expression of BAIAP3 in LA patients. The mRNA level of BAIAP3 was determined using real-time PCR in whole blood from type I LA subjects (n=12) and controls (n=12). (D) Diagram of predicted miR-1972 binding sites on the 3’UTR of BAIAP3 gene. The 3’UTR mutant sequence of BAIAP3 inserted in dual-luciferase reporter genes is indicated in blue. (E) Effect of hsa-miR-1972 on BAIAP3 mRNA transcription in 293T cells using the dual-luciferase reporter assay. Relative luciferase activity was analyzed in 293T cells co-transfected with pMRI-REPORT vector carrying either wild-type or mutated 3’UTR of BAIAP3 and hsa-miR-1972 mimic or relative negative control. (F) Effect of hsa-miR-1972 on the expression level of BAIAP3 protein in 293T cells observed in western blots. 293T cell lines were transfected with miR-1972 mimic (125 nM), miR-1972 inhibitor (125 nM), negative mimic control (125 nM), or negative inhibitor control (125 nM). Protein levels of BAIAP3 were detected at 48 h after post-transfection using western blotting. Tubulin was used as an internal control. (G) Quantitative analysis of results obtained from three independent experiments which are presented in (F). The results show that hsa-miR-1972 can significantly inhibit the expression of BAIAP3 in vitro. *P < 0.05, *P < 0.01, *P < 0.001.