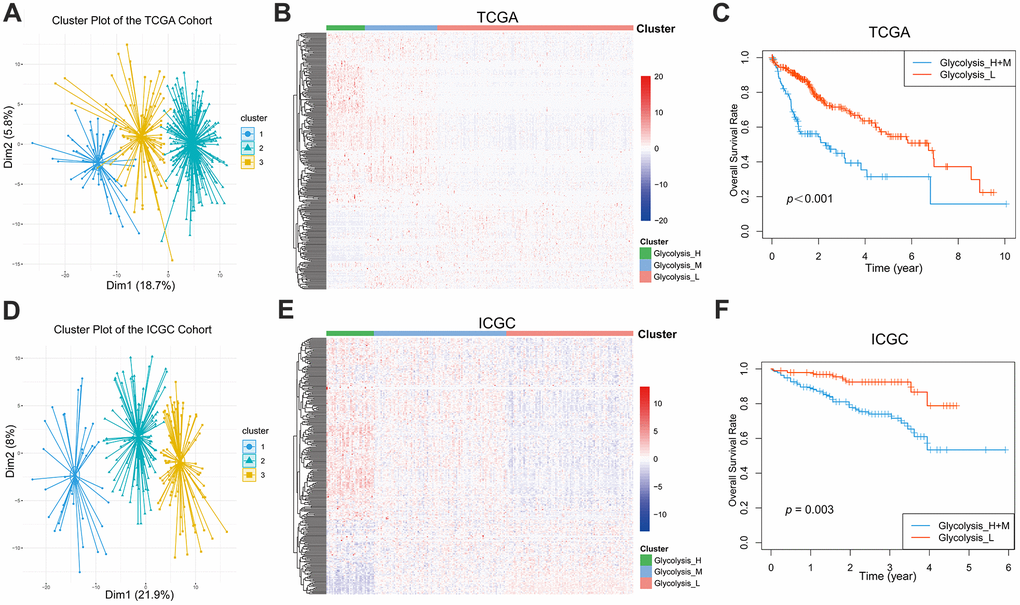
Figure 1.Glycolysis-associated genes identified distinct HCC clusters with different OS. (A, B) Three distinct clusters were generated by hierarchical clustering analysis based on the expression level of the 288 glycolysis-associated genes in the TCGA. (D, E) Three glycolysis-associated clusters were generated in the ICGC. (C, F) Kaplan-Meier survival curves of different glycolysis subtypes in the TCGA and ICGC. The Glycolysis-H+M subgroup had a worse OS than the Glycolysis-L subgroup (TCGA cohort: log-rank p<0.001; ICGC cohort: log-rank p= 0.003). HCC, hepatocellular carcinoma; OS, overall survival.