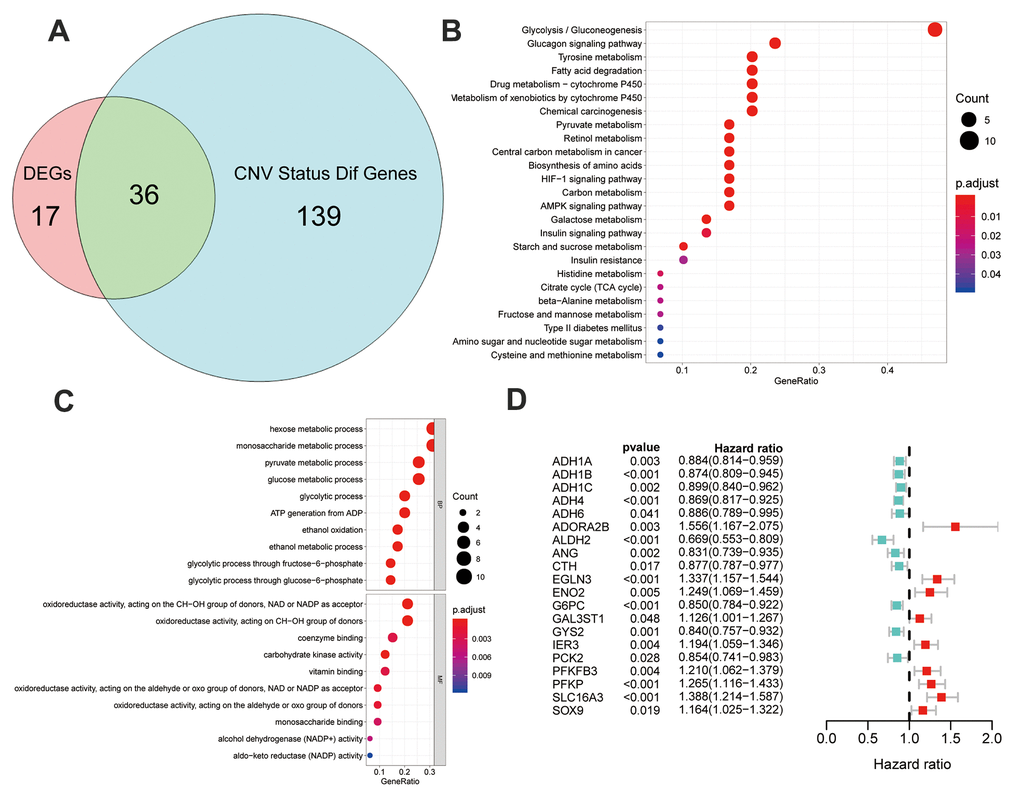
Figure 4.Identification and analysis of different status genes among glycolysis subgroups based on multi-omics data. (A) Venn diagram showed 36 overlapped genes in DEGs and genes with differently CNV status. (B) The results of gene ontology functions analysis of the 36 multi-omics based different status glycolysis genes. (C) KEGG pathways enrichment analysis of the 36 multi-omics based different status glycolysis genes. (D) 20 MOG-DEGs with a significant correlation with OS by univariable Cox analyses in the TCGA.