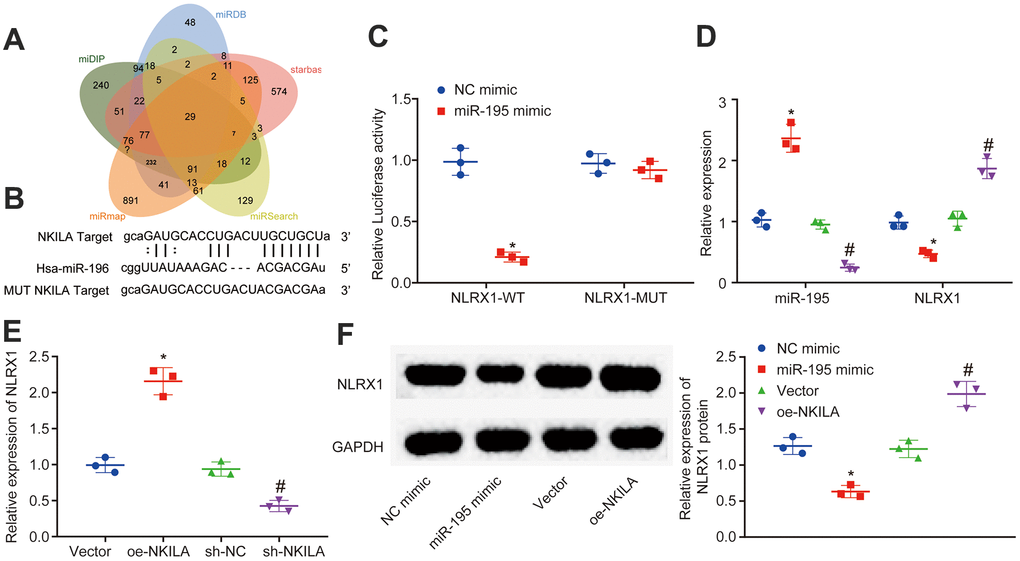
Figure 4.NKILA acts as ceRNA of miR-195 to upregulate NLRX1. (A) Venn analysis of target genes of miR-195 obtained from miDIP, miRDB and starbase. (B) putative binding sites between NLRX1 and miR-195 predicted in RNA22. (C) the binding relationship between NLRX1 and miR-195 verified by dual-luciferase reporter gene assay. * p < 0.05 compared with the treatment of NC mimic. (D) NLRX1 and miR-195 expression in neurons after alteration of miR-195 detected by RT-qPCR. * p < 0.05 compared with the treatment of NC mimic. # p < 0.05 compared with the treatment of NC inhibitor. (E) NLRX1 mRNA expression in neurons after alteration of NKILA detected by RT-qPCR. * p < 0.05 compared with the treatment of vector. # p < 0.05 compared with the treatment of sh-NC. (F) NLRX1 protein expression in neurons after overexpression of NKILA or miR-195 measured by Western blot analysis. * p < 0.05 compared with the treatment of NC mimic. # p < 0.05 compared with the treatment of vector. All data were measurement data and expressed as mean ± standard deviation. Unpaired t test was used for comparison between two groups. The one-way ANOVA was used for comparison among multiple groups, followed by Tukey’s post-hoc test. The cell experiment was repeated three times.