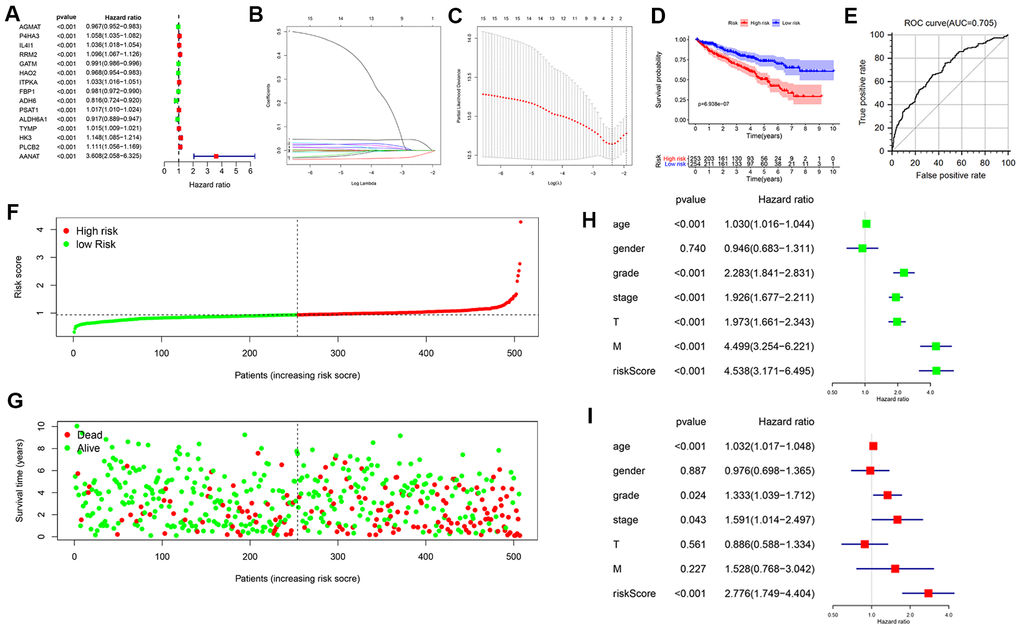
Figure 2.Construction of a prognostic MRG signature based on TCGA-KIRC cohort. (A) Identification of 15 MRGs in significant association with OS by univariate Cox regression analysis. (B, C) Screening of candidate MRGs used for the construction of the predictive signature using LASSO regression analysis. (D) Survival curves of KIRC patients assigned to high-and low-risk groups based on individual risk scores derived from the prognostic signature. (E) ROC analysis demonstrating survival prediction accuracy. (F) Distribution of risk scores. (G) Survival status for KIRC patients in the high- and low-risk groups. Univariate (H, I) multivariate Cox regression analysis were used to verify that the metabolism-related prognostic signature represents an independent prognostic factor for KIRC patients.