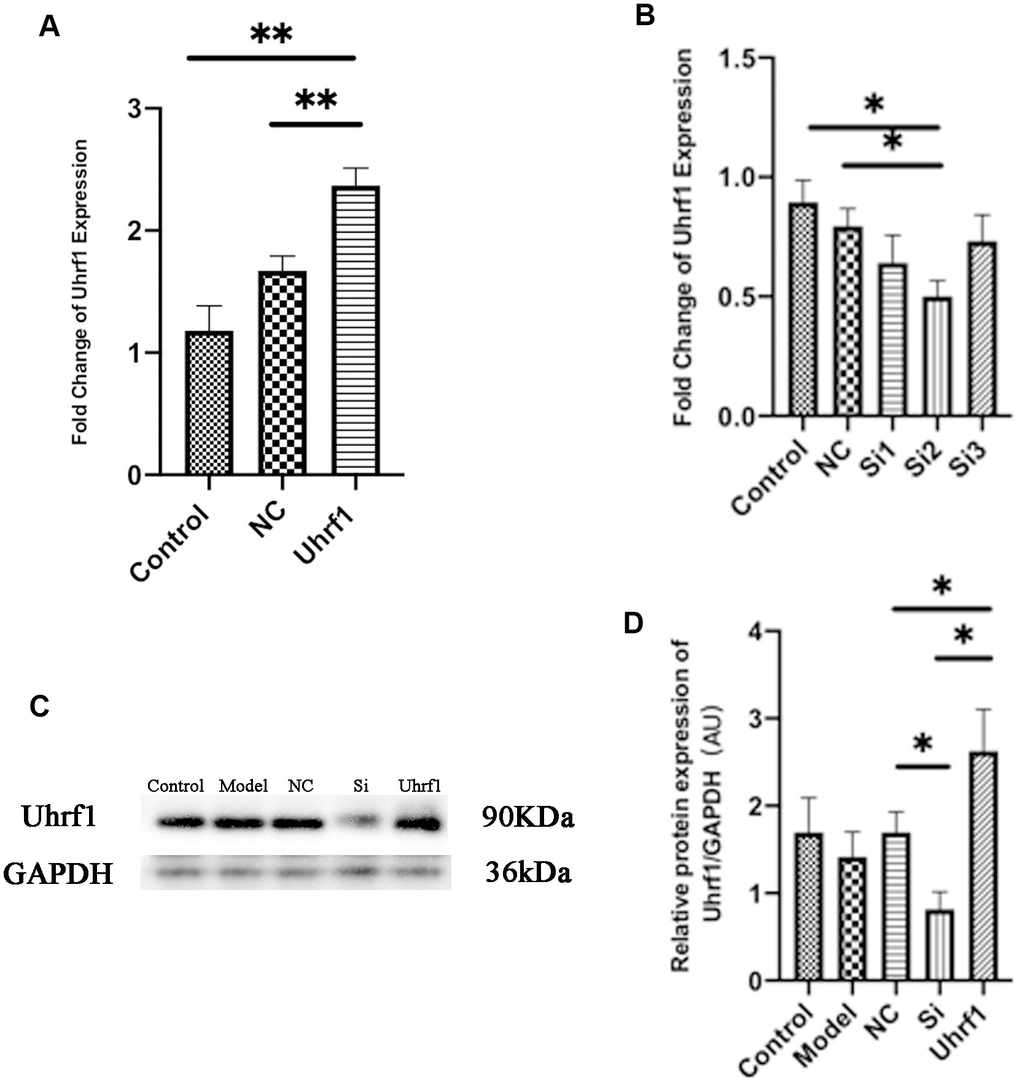
Figure 2.Detection of transfection efficiency of overexpression and knockdown of Uhrf1 plasmid. (A) The relative mRNA expressions of Uhrf1 in Myocardial ischemia-reperfusion model in vitro with Uhrf1 plasmid were determined by qRT-PCR. NC, negative control of plasmid transfection; Uhrf1, Uhrf1 overexpression. (B) The relative mRNA expressions of Uhrf1 in Myocardial ischemia-reperfusion model in vitro with Uhrf1 siRNA plasmid were determined by qRT-PCR. NC, negative control of RNAi; si1-3, siRNA targeting different mRNA regions. (C) Western blot was used to detect the expression level of Uhrf1 protein in each group. GAPDH serves as a loading control. Model, in vitro oxidative stress model; NC, negative control of RNAi; si, RNAi knockdown of Uhrf1; Uhrf1, Uhrf1 overexpression. (D) Expression of Uhrf1 protein relative to GAPDH data from 3 biological repeats is shown. Data shown are mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. N=3 per group.