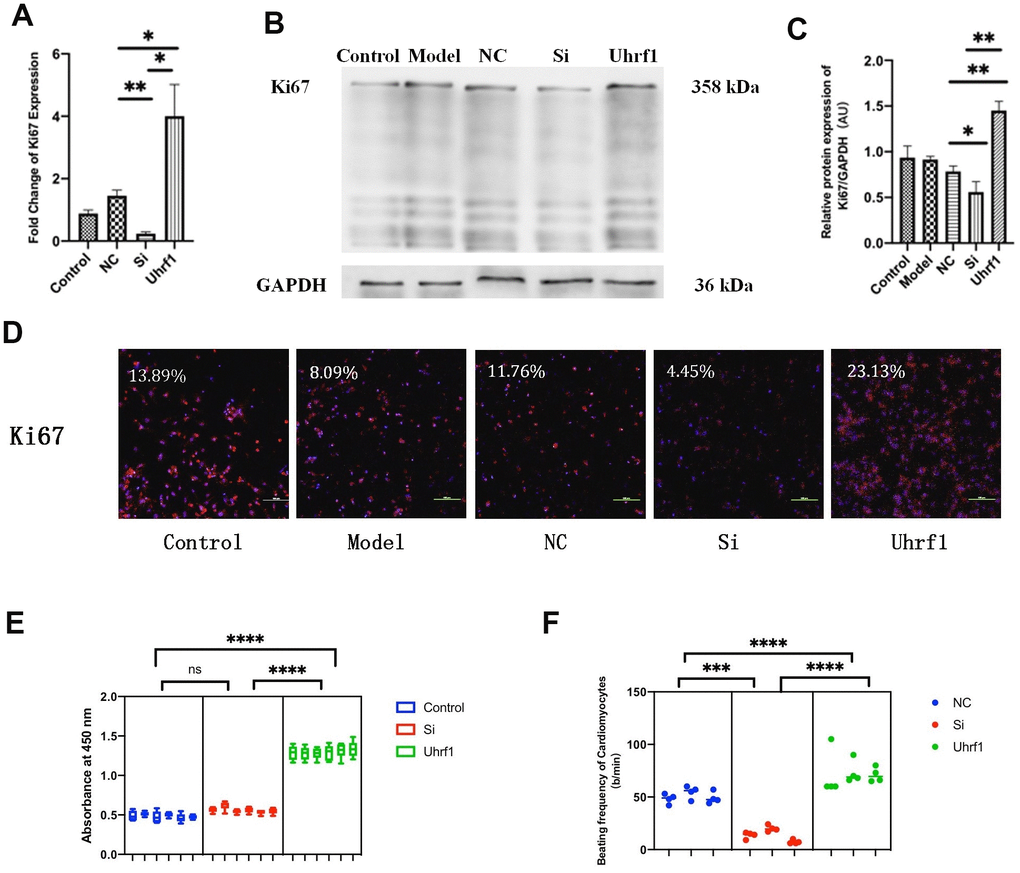
Figure 4.The effect of Uhrf1 on cell cycle and viability in myocardial ischemia-reperfusion injury. (A) The relative mRNA expressions of Ki67 in Myocardial ischemia-reperfusion model in vitro were determined by qRT-PCR. (B) Western blot was used to detect the expression level of Ki67 protein in each group. GAPDH serves as a loading control. (C) Expression of Ki67 protein relative to GAPDH data from 3 biological repeats is shown. (D) Detection of Ki67 protein expression and semi quantitative analysis by immunofluorescence microscopy (x200). Scale bar, 100μm. (E) The cardiomyocytes viability in knockdown and overexpression Uhrf1 groups was detected by CCK8. (F) The beating times per minute of cardiomyocytes in overexpression and knockdown group. Data shown are mean ± SD. *P <0.05, **P <0.01, ***P <0.001, ****P <0.0001. N=3 per group. Model, in vitro oxidative stress model; NC, negative control of RNAi; si, RNAi knockdown of Uhrf1; Uhrf1, Uhrf1 overexpression.