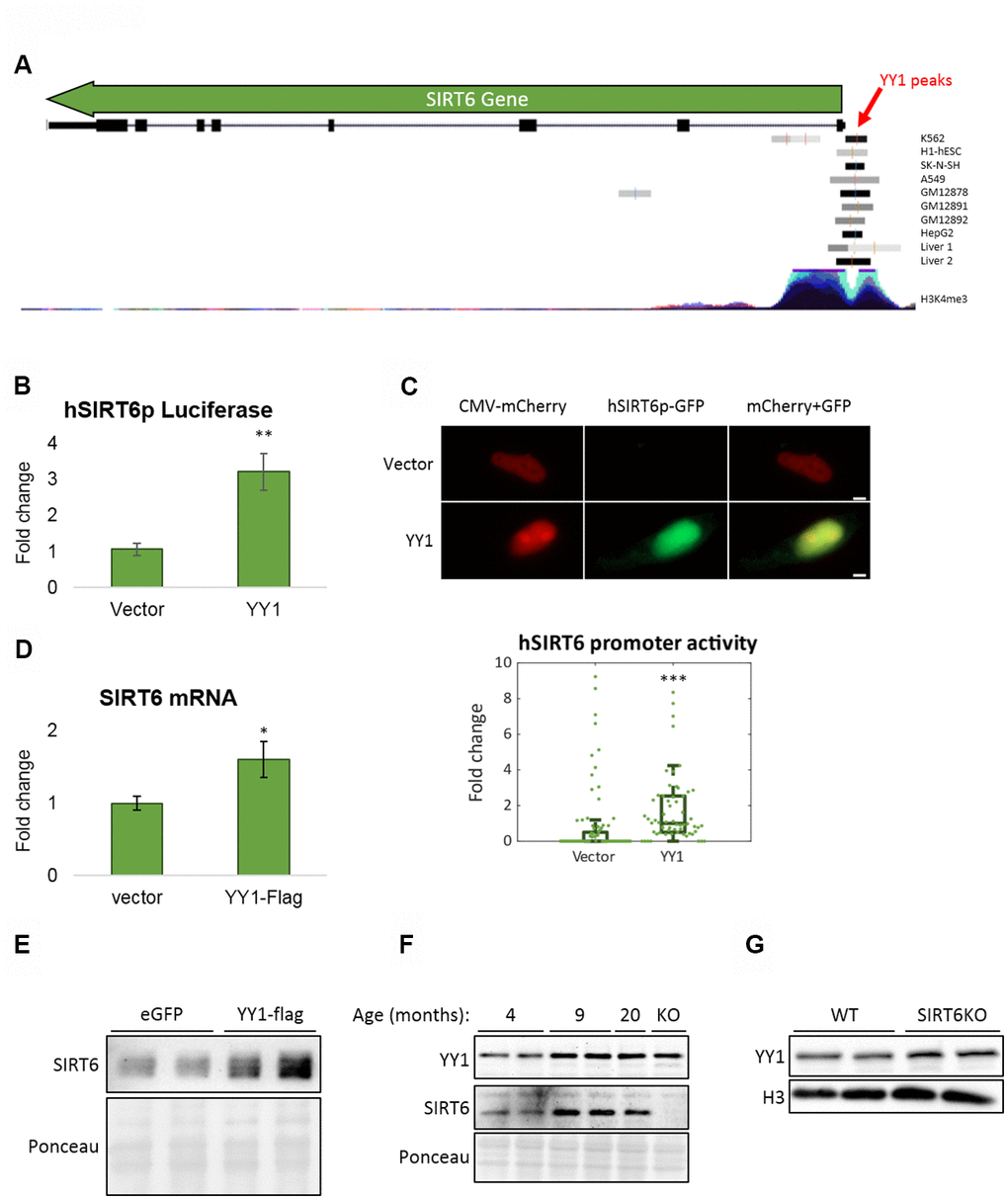
Figure 5.YY1 regulates SIRT6 promoter. (A) YY1 ChIP-seq data in SIRT6 locus. SIRT6 locus is marked with a green arrow; YY1 peaks are marked in black or grey shades; tested cell lines are marked on the right side of each YY1 peak. (B) hSIRT6p-Firefly Luciferase assay with an empty vector or YY1-Flag overexpression vector. Chosen promoter region – 1000 bases before TSS and 2000 bases after (including the first 2 exons and the intron in between). (C) Human SIRT6 promoter activation tested using a hSIRT6p-GFP-CMV-mCherry vector, co-transfected with either empty or YY1-overexpressing vector. Upper panel – representative cell images; lower panel – an unbiased quantification of GFP/mCherry ratio per cell. Box plots represent quartile range, whiskers represent 10% and 90% of datapoints, horizontal line represents median. (D) SIRT6 mRNA levels in cells transfected with either an empty or YY1-flag overexpression vector. (E) SIRT6 protein levels in cells transfected with either eGFP- or YY1-overexpressing vectors. (F) Brains of WT mice at different ages and SIRT6-KO mouse (serves as a control for SIRT6 specificity). (G) Protein blots of WT or SIRT6-KO SH-SY5Y cells. * - p-value < 0.05, ** - p-value < 0.01, *** - p-value < 0.001.