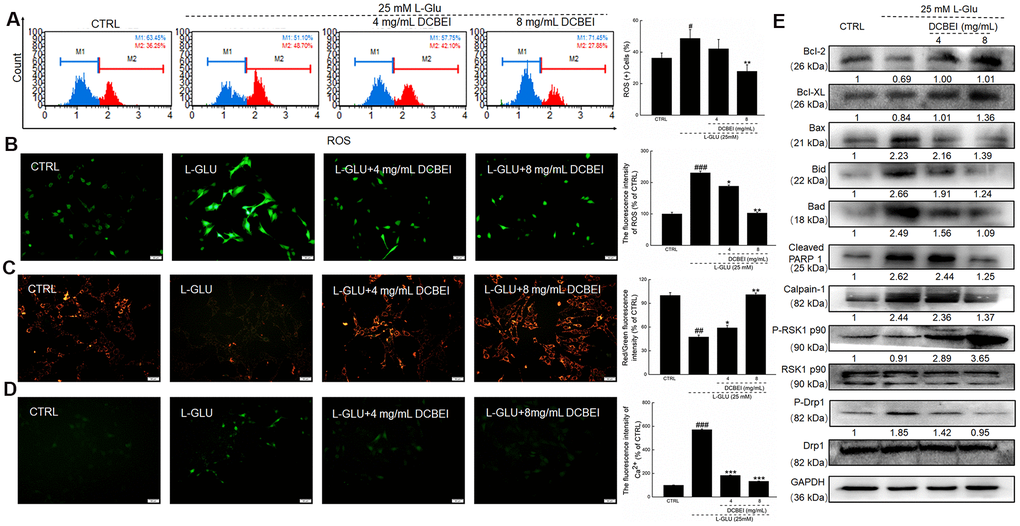
Figure 2.HT22 cells were pre-incubated with DCBEI (4 mg/mL or 8 mg/mL) for 3 h, and co-treated with L-Glu for a further 24 h. (A) DCBEI reduced the intracellular ROS activity in L-Glu-treated HT22 cells (n = 3). (B) DCBEI reduced L-Glu-induced ROS production (200×) (Scale bar: 50 μm) (n = 3). (C) DCBEI attenuated L-Glu-induced MMP dissipation (200×) (Scale bar: 50 μm) (n = 3). (D) Fluo 4-AM staining indicated that DCBEI suppressed L-Glu-induced Ca2+ increases (200×) (Scale bar: 50 μm) (n = 3). (E) DCBEI regulated the expression of proteins related to mitochondrial function. DCBEI increased the levels of anti-apoptotic proteins (Bcl-2, Bcl-xL and P-RSK1 p90) and reduced the levels of pro-apoptotic proteins (Bax, Bid, Bad, cleaved PARP-1, Calpain-1 and p-Drp1) in L-Glu-treated HT22 cells. Quantification data were normalized to GAPDH and the corresponding total proteins, and reported as fold change relative to CTRL (n = 3). Data are expressed as a percentage of corresponding control cells and means ± S.D. # P < 0.05, ## P < 0.01 and ### P < 0.001 vs. CTRL, * P < 0.05, ** P < 0.01 and *** P < 0.001 vs. L-Glu-treated cells.