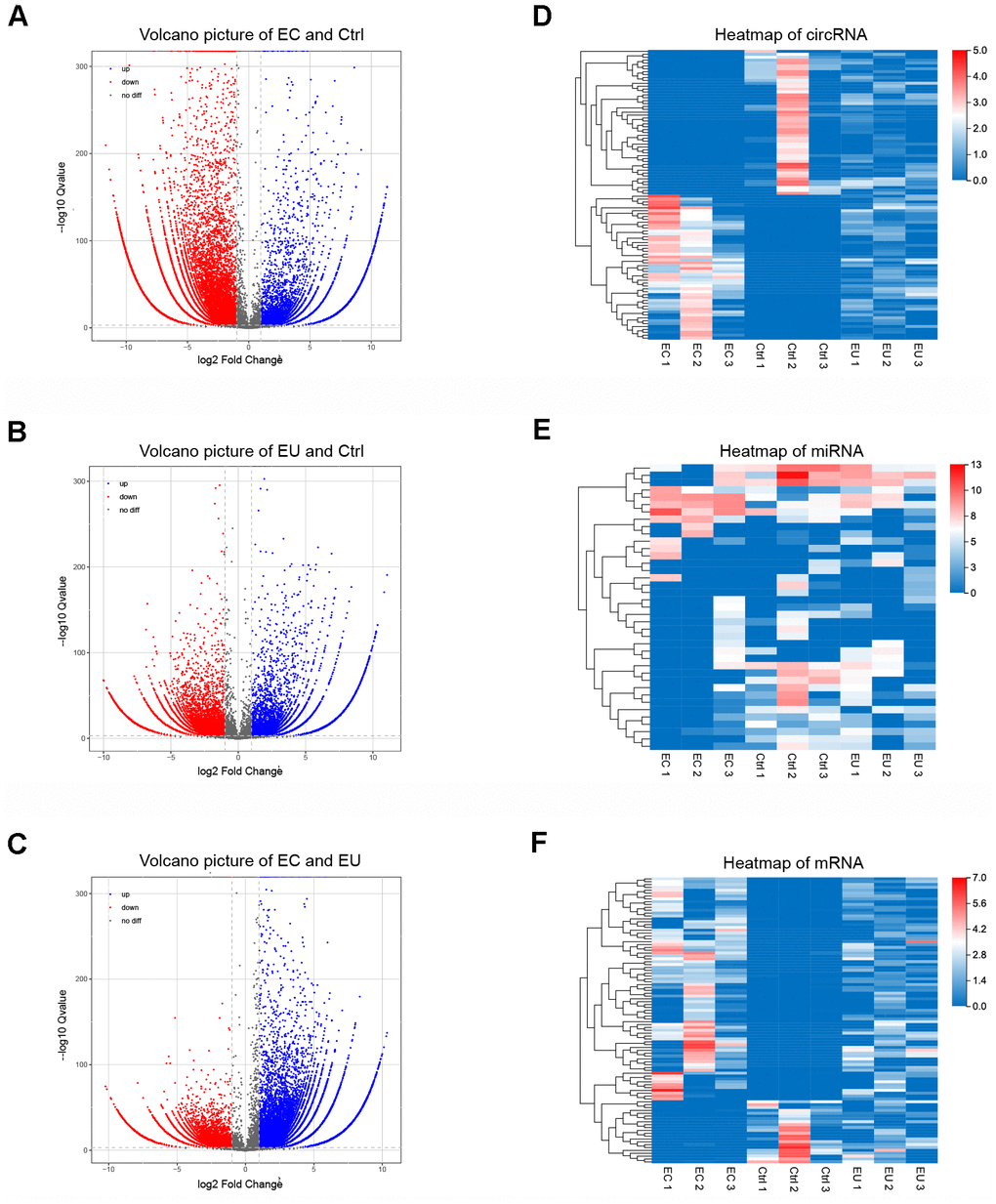
Figure 2.Identification of exosomal DECs, DEMis and DEMs in endometriosis. (A–C) Volcano plots of exosomal DECs based on the |log fold-change| between the EC and Ctrl groups (A), the EU and Ctrl groups (B) and the EC and EU groups (C). (D–F) Heatmap showing the expression of overlapping DEGs in the EC vs. Ctrl, EU vs. Ctrl and EC vs. EU comparisons, including the top 100 DECs, top 100 DEMis and top 100 DEMs. DECs, DEMis and DEMs: Differentially expressed circRNAs, miRNAs and mRNAs.