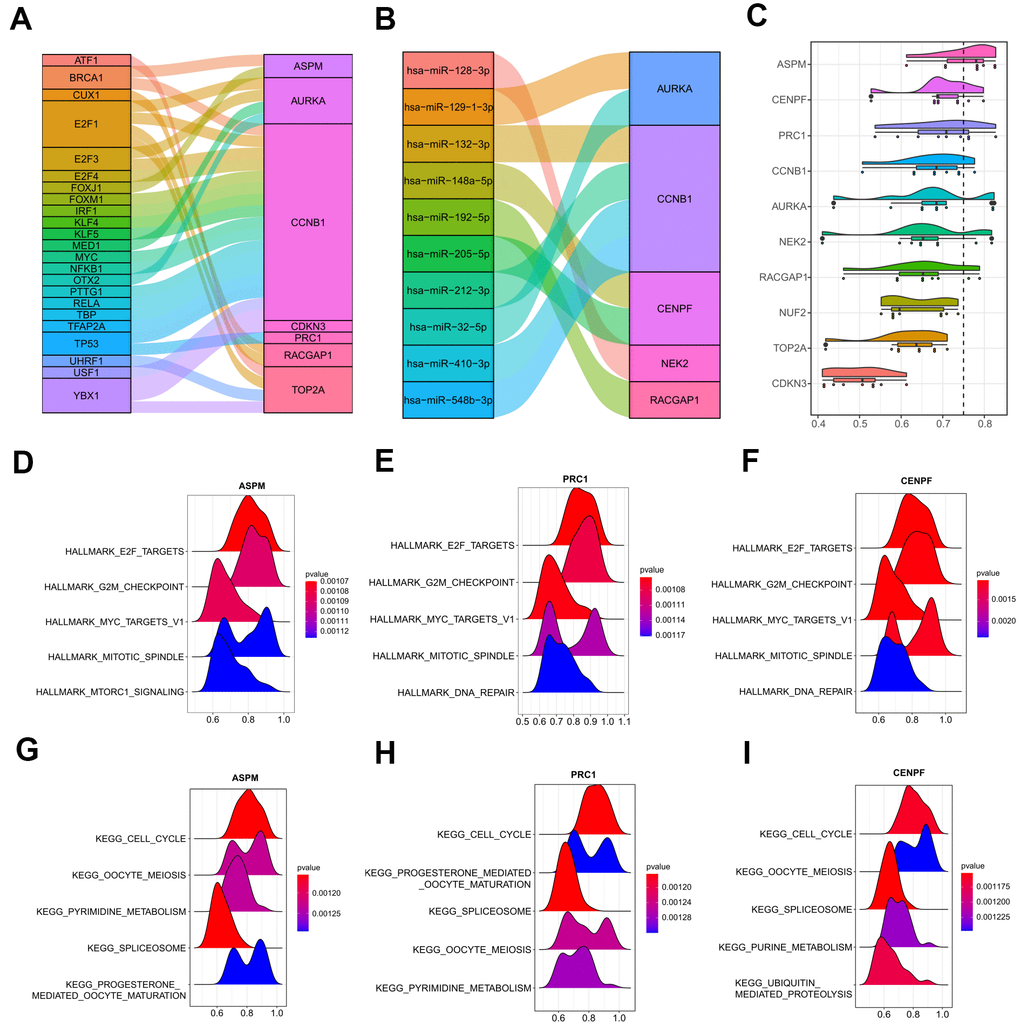
Figure 11.Upstream regulations of the ten hub genes and GO semantic similarities analysis. (A) The transcription factor-hub gene network predicted by miRNet. (B) 10 function MTIs predicted through miRTarBase 8.0. (C) Raincloud plot showing the ranking list of function semantic similarities for the 10 hub genes using the ICGC-LIRI-JP dataset. ASPM, CENPF, and PRC1 were the top three hub genes with the highest scores. (D–F) GSEA results of ASPM, CENPF, and PRC1 based on the hallmark gene set. (G–I) GSEA results of ASPM, CENPF, and PRC1 based on the KEGG database. GO, gene ontology. MTIs, miRNA-target interactions. KEGG, Kyoto Encyclopedia of Genes and Genomes.