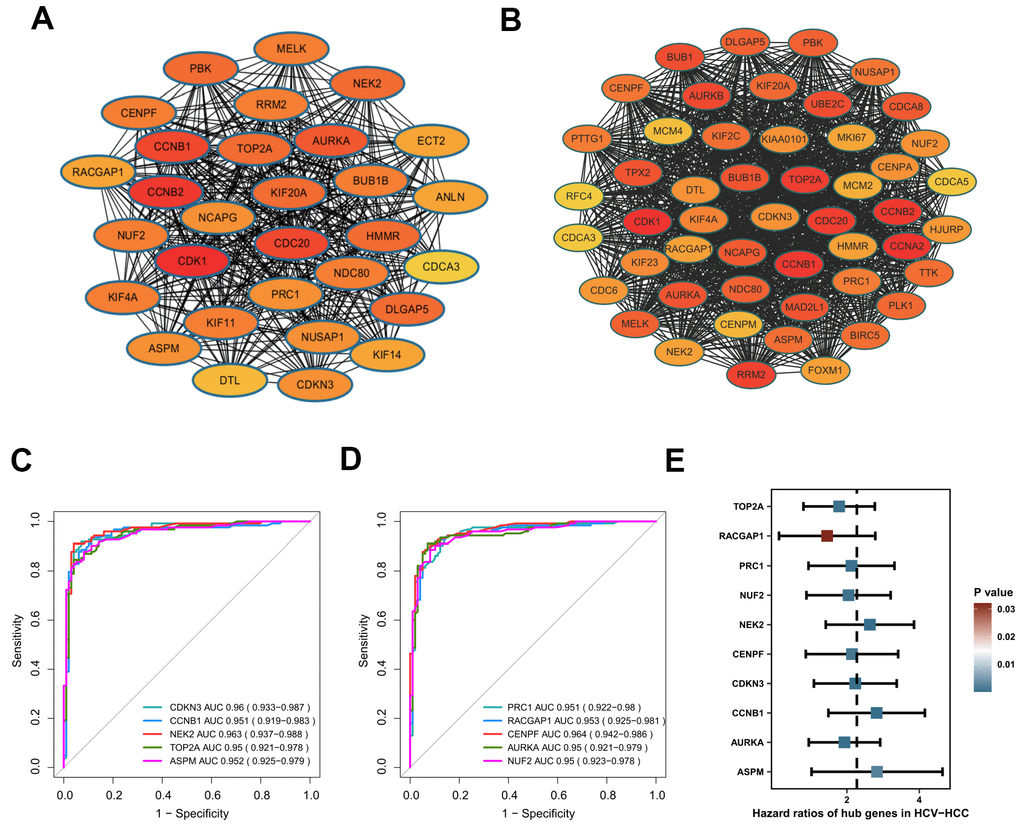
Figure 4.Identification of hub genes in HCV-HCC. (A) The most significant cluster identified from the DEGs-PPI network. (B) The WGCNA-PPI network constructed by the turquoise module. (C, D) ROC curves showing the AUROC scores and AUC (95%CI) of the 10 hub genes for discriminating tumor from normal samples based on the ICGC-LIRI-JP dataset. Colored lines indicate the ROC curve for each hub gene, and the grey line indicates the reference line. (E) Forest plot presenting the results of the univariate Cox regression analysis for the 10 hub genes. HCV-HCC, HCV- associated HCC. DEGs, differentially expressed genes. PPI, protein-protein interaction. WGCNA, Weight Gene Co-expression Network Analysis. ROC, receiver operating characteristic. 95%CI, 95% confidence interval.