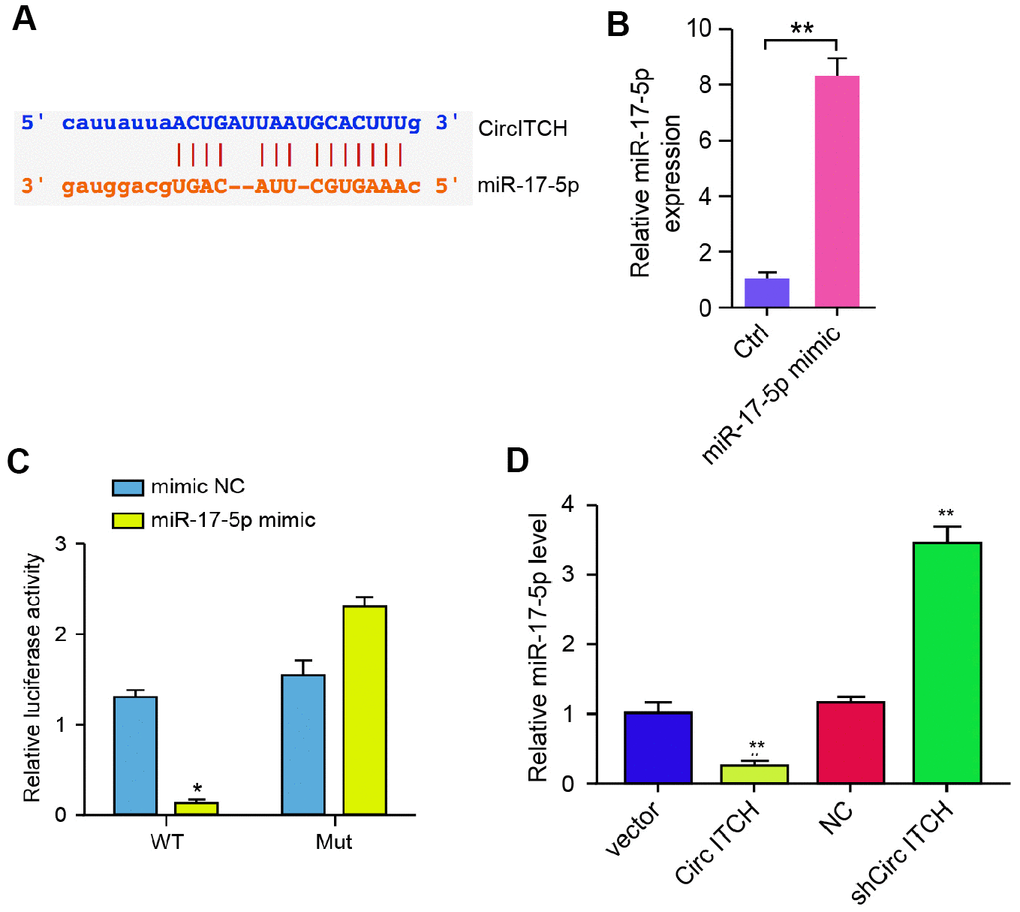
Figure 3.CircITCH serves as a miR-17-5p sponge in NP cells. (A) The potential interaction between circITCH and miR-17-5p was identified by the bioinformatic analysis using ENCORI (http://starbase.sysu.edu.cn/index.php). (B, C) The NP cells were treated with the miR-17-5p mimic or control mimic. (B) The expression levels of miR-17-5p were measured by qPCR in the cells. (C) The luciferase activities of wild type circITCH (circITCH WT) and circITCH with the miR-17-5p-binding site mutant (circITCH MUT) were determined by luciferase reporter gene assays in the cells. (D) The NP cells were infected with lentiviral plasmids carrying circITCH shRNA or corresponding control shRNA or transfected with the pcDNA3.1 or the pcDNA3.1-circITCH overexpression vector. The expression of miR-17-5p was analyzed by qPCR in the cells. Data are presented as mean ± SD. Statistic significant differences were indicated: * P < 0.05, ** P < 0.01.