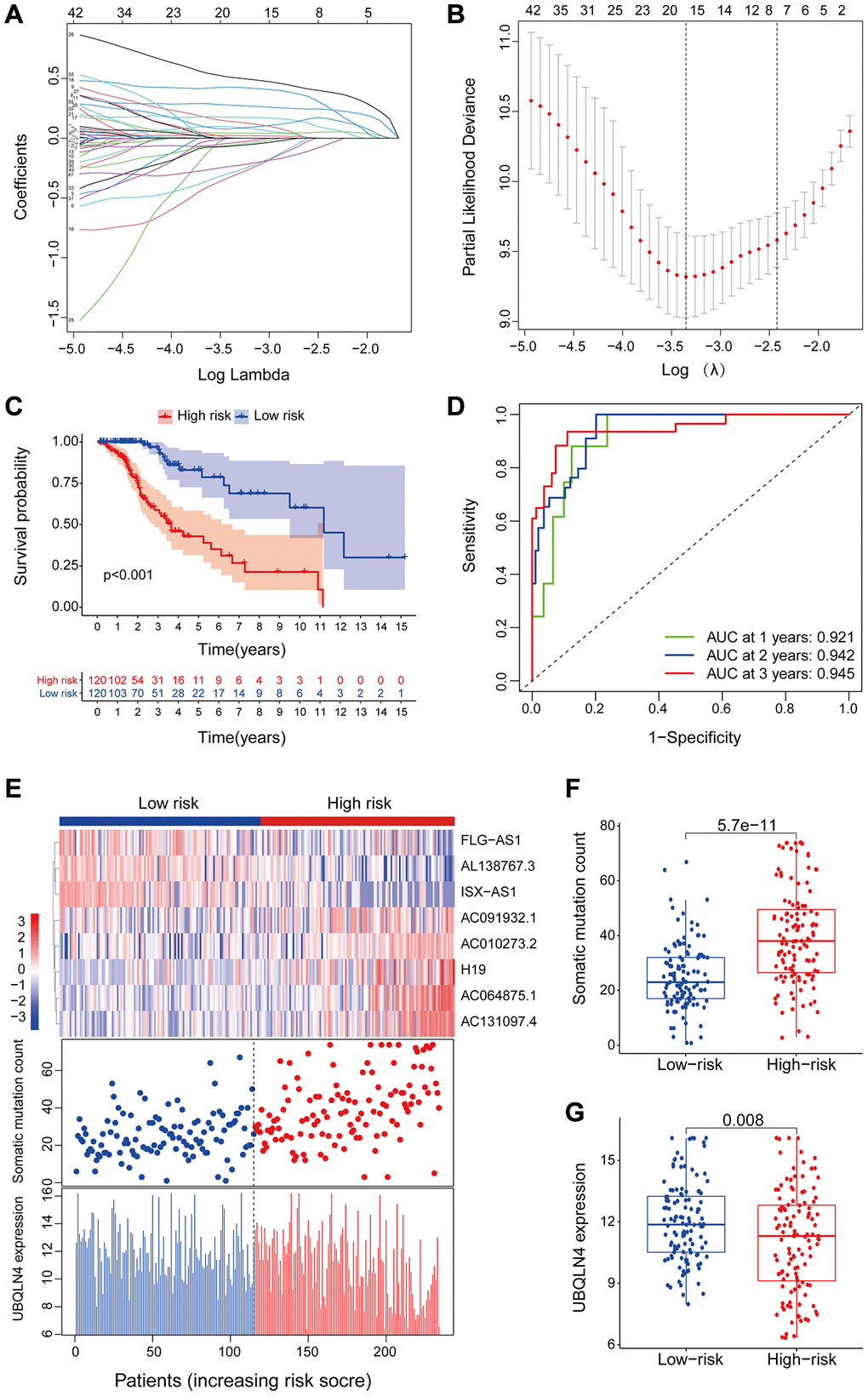
Figure 3.LncRNAs signature of the genomic instability used to predict outcomes in the training set. (A) Lasso Cox analysis identified 16 lncRNAs associated with genomic instability that were highly associated with prognosis. (B) Determination of the optimal value of penalty parameters through 1000 replicates of cross-validation. (C) Kaplan-Meier estimation of GILncSig-predicted overall survival of low- or high-risk patients in the training set. (D) Time-dependent ROC curves of GILncSig at 1, 2 and 3 years. (E) Distribution of cumulative somatic mutations and expression of UBQLN4 in high- and low-risk groups in the GILncSig model of low-grade glioma patients. (F) Box plot for distribution of cumulative somatic mutations in the low- and high-risk groups of LGG patients. (G) Box plots for UBQLN4 gene expression in low- and high-risk groups of LGG patients.