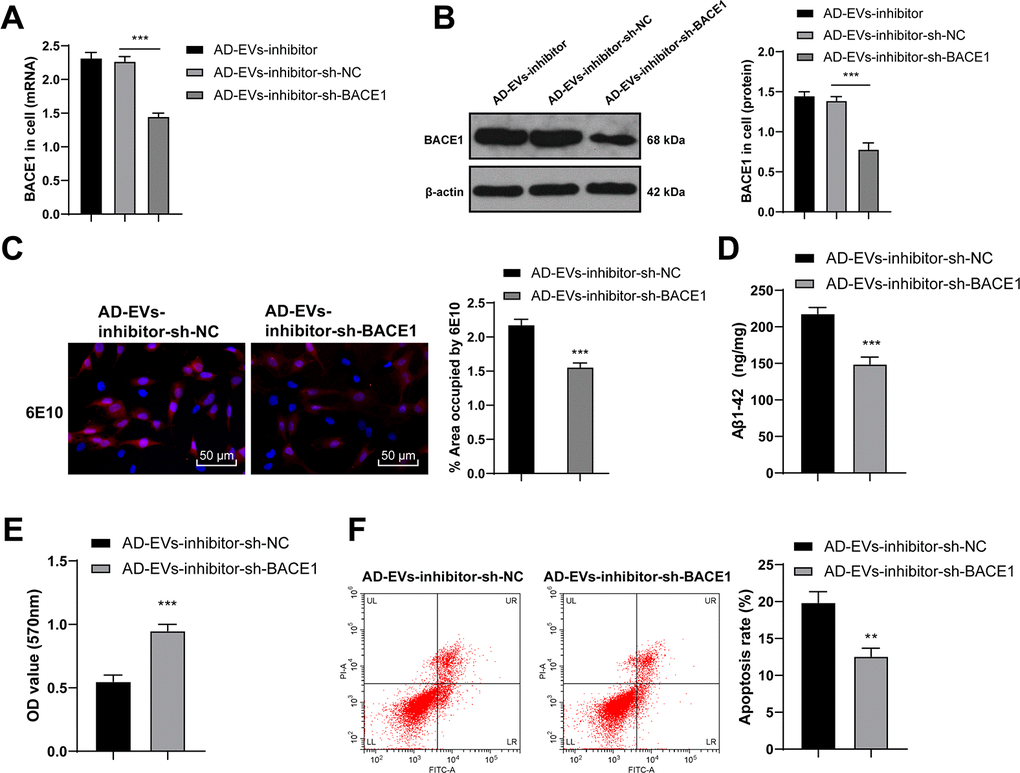
Figure 6.BACE1 knockdown reverses the effects of EVs-inhibitor on AD. AD-EVs-inhibitor-treated neurons were transfected with BACE1 shRNA (AD-EVs-inhibitor-sh-BACE1), with AD-EVs-inhibitor-treated neurons transfected with sh-NC (AD-EVs-inhibitor-sh-NC) as control. (A, B) RT-qPCR and WB were used to detect the silencing effect of shRNA on BACE1; (C) Immunofluorescence assay was used to detect the content of Aβ in AD hippocampal neurons; (D) ELISA was used to detect level of Aβ1-42; (E) MTT assay was used to detect the viability of AD hippocampal neurons; (F) Flow cytometry was used to detect the apoptosis rate of AD hippocampal neurons. The experiment was repeated three times, and the data were expressed as mean ± standard deviation. Comparisons between two groups were analyzed using independent sample t-test; comparisons among multiple groups were analyzed using one-way ANOVA, followed by Tukey’s multiple comparisons test or Sidak’s multiple comparisons test. ***p < 0.001.