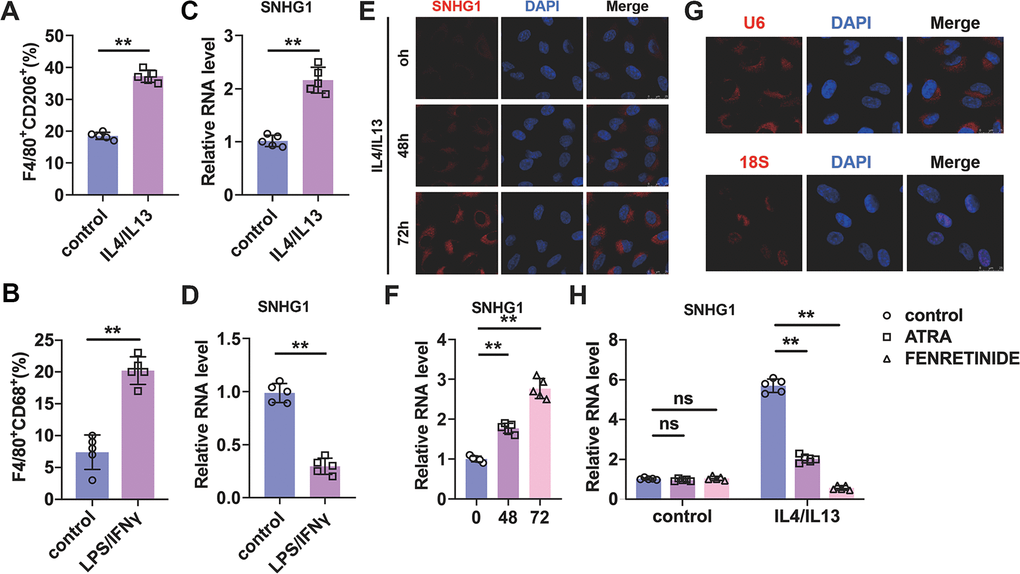
Figure 1.Identification of lncRNA-SNHG1 as a potential mediator of M2 macrophage polarization. (A, B) Flow-cytometric analysis was performed to analyze the percentage of F4/80+CD206+ or F4/80+CD86+ cells in BMDMs. (C, D) qRT-PCR analysis was performed to analyze the RNA level of lncRNA-SNHG1 in BMDMs. BMDMs were treated with IL4/IL13 or LPS/INFγ for 72h. Two-tailed t-test was used for the statistical analysis. n=5 independent cell cultures. The bar indicates the SD values. **P<0.01. (E) The FISH assay was conducted to detect the expression of lncRNA-SNHG1. RAW264.7 cells were treated with IL4/IL13 for 72h. (F) Quantitative analysis of E. Two-tailed t-test was used for the statistical analysis. n=5 independent cell cultures. The bar indicates the SD values. **P<0.01. (G) The localization of U6 snRNA and 18S rRNA was tested by FISH assay in RAW264.7 cells. (H) Two inhibitors of M2 macrophage polarization were used to block M2 polarization of RAW264.7 cells. qRT-PCR analysis was performed to analyze the RNA level of lncRNA-SNHG1. Two-tailed t-test was used for the statistical analysis. n=5 independent cell cultures. The bar indicates the SD values. **P<0.01.