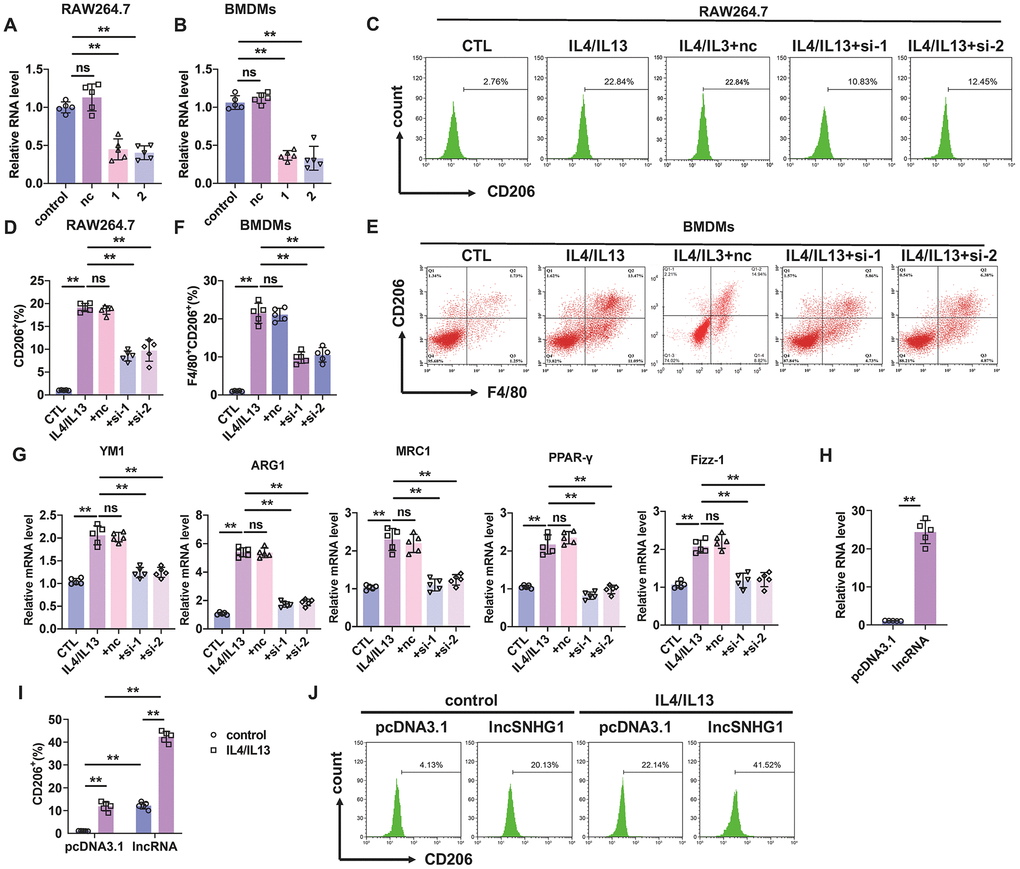
Figure 2.Silencing of lncRNA-SNHG1 inhibited M2 macrophage polarization. (A, B) Two siRNAs of lncRNA-SNHG1 were construct and knockdown efficiency was confirmed in RAW264.7 cells and BMDMs using qRT-PCR analysis. Two-tailed t-test was used for the statistical analysis. n=5 independent cell cultures. The bar indicates the SD values. **P<0.01. (C, D) Flow-cytometric analysis was performed to analyze the percentage of CD206+ cells. RAW264.7 cells were transfected with two siRNAs of lncRNA-SNHG1 and negative control siRNA for 48h and treated with IL4/IL13 or LPS/INFγ for 72h. one-way ANOVA was used for the statistical analysis. n=5 independent cell cultures. The bar indicates the SD values. **P<0.01. (E, F) Flow-cytometric analysis was performed to analyze the percentage of F4/80+CD206+ cells. BMDMs were transfected with two siRNAs of lncRNA-SNHG1 or negative control siRNA for 48h and treated with IL4/IL13 or LPS/INFγ for 72h. one-way ANOVA was used for the statistical analysis. n=5 independent cell cultures. The bar indicates the SD values. **P<0.01. (G) mRNA of M2 polarization marker genes in RAW264.7 cells was quantified by qRT-PCR analysis. one-way ANOVA was used for the statistical analysis. n=5 independent cell cultures. The bar indicates the SD values. **P<0.01. (H) Overexpression of lncRNA-SNHG1 was identified via performing qRT-PCR analysis. pcDNA3.1 empty vector acted as a negative control. one-way ANOVA was used for the statistical analysis. n=5 independent cell cultures. The bar indicates the SD values. **P<0.01. (I, J) Flow-cytometric analysis was performed to analyze the percentage of CD206+ cells. RAW264.7 cells were transfected with expression plasmid of lncRNA-SNHG1 or pcDNA3.1 and treated with IL4/IL13 or LPS/INFγ for 72h. one-way ANOVA was used for the statistical analysis. n=5 independent cell cultures. The bar indicates the SD values. **P<0.01.