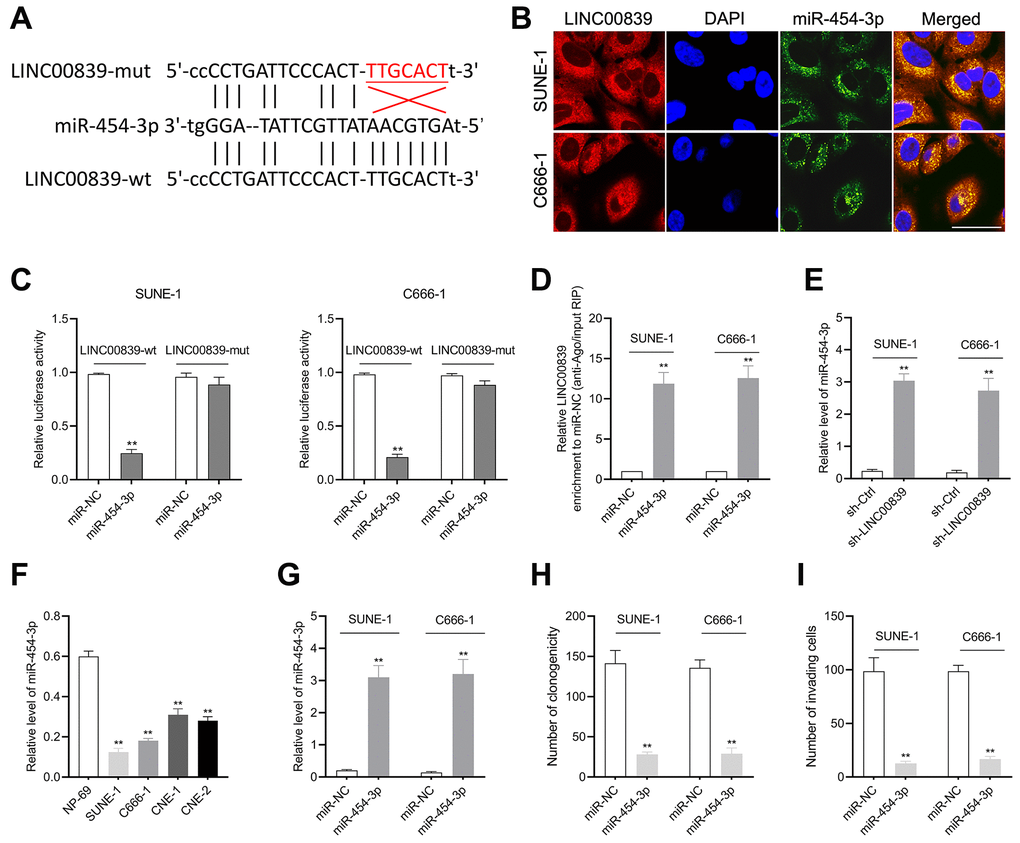
Figure 3.The relationship between LINC00839 and miR-454-3p in SUNE-1 and C666-1 cells. (A) Online software starBase showed the sequence alignment of miR-454-3p with the putative binding sites within LINC00839. (B) FISH analysis of the cellular colocalization of LINC00839 and miR-454-3p in SUNE-1 and C666-1 cell. Nuclei were stained with DAPI (scale bar, 20 μm). (C) The luciferase reporter assay demonstrated the influence of miR-454-3p on the luciferase activity in C666-1 and SUNE-1 cells transfected with LINC00839-wt or LINC00839-mut vector. (D) RIP assay was performed to further identify the potential binding of LINC00839 and miR-454-3p. (E) Levels of mIR-454-3p were detected by qPCR after cells transfected with sh-LINC00839. (F) qPCR examined miR-454-3p level in NPC cell lines (SUNE-1, CNE-1, C666-1 and CNE-2) compared with that in NP-69. **P < 0.01 compared with NP-69. (G) qPCR analysis testified miR-454-3p expression in C666-1 and SUNE-1 cells after transfected with miR-NC or miR-454-3p mimic. (H) After transfection, the growth of NPC cells was determined by colony formation assay. (I) The invasion of NPC cells was analyzed using Transwell assay. **P < 0.01 compared with miR-NC group.